15 ggplot2
1 Data vizualization with ggplot2
1.1 Why ggplot2?
-
base Rplot functions require more effort and expertise to create high-quality publishable graphs - Over the years, many R packages (e.g.
lattice,grid, etc.) were introduced to overcome the limitations ofbase Rplot functions - The newest addition to R plot functions is
ggplot2package and it can be used to produce elegant plots without much effort!
1.2 The Grammar of Graphics
- ggplot2 is built on the grammar of graphics, a structured system for describing graphs
- Every plot is created from the same components:
- data
- aesthetic mappings
- geometric objects (geoms)
- optional: facets, scales, themes, statistics
- To load
ggplot2package to the current R environment
2 Scatter Plot: The Basic Structure
ggplot(data = penguins) +
geom_point(mapping = aes(x = bill_length_mm, y = flipper_length_mm))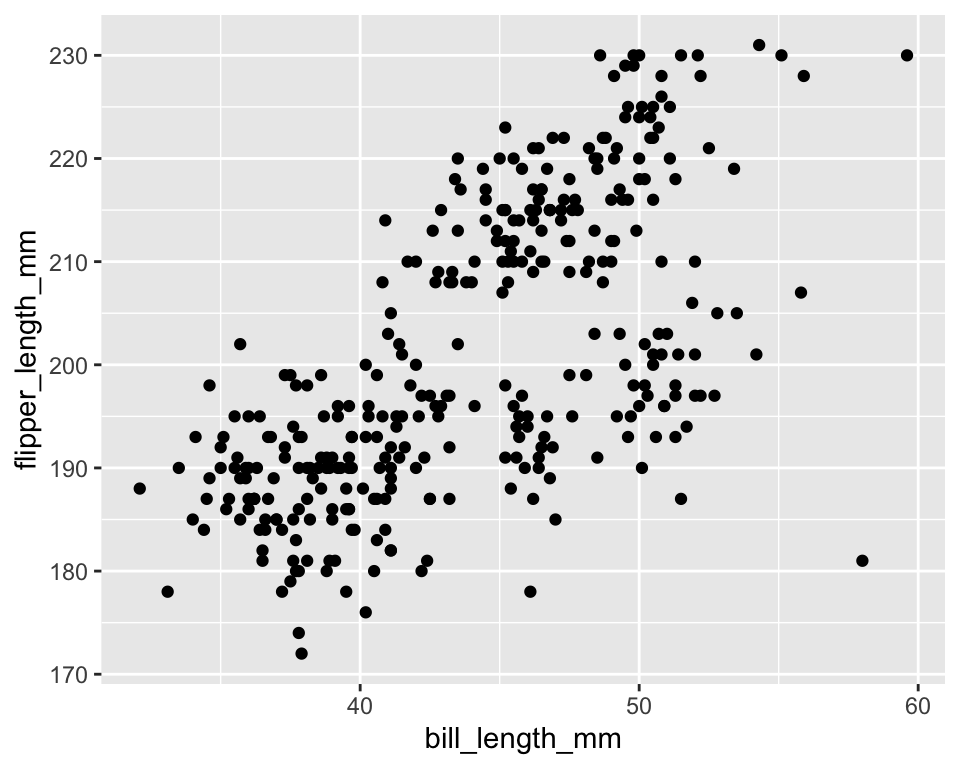
2.1 How a ggplot is built
ggplot(data = penguins) +
geom_point(mapping = aes(x = bill_length_mm, y = flipper_length_mm))ggplot(data = penguins)
- Creates an empty coordinate system to which multiple layers can be added.
- All ggplot2 visualizations begin with the
ggplot()function. - The argument
dataspecifies the data frame used for the plot. - Additional layers are added using the plus sign.
geom_point
- Geometric objects (called
geom) are the shapes we put on a plot (e.g. points, bars, etc.). - You can have an unlimited number of layers, but at a minimum a plot must have at least one geom
-
geom_point()makes a scatter plot by adding a layer of points. -
geom_line()adds a layer of lines connecting data points. -
geom_col()adds bars for bar charts. -
geom_histogram()makes a histogram. -
geom_boxplot()adds boxes for boxplots
-
mapping = aes()
- Each type of
geomusually has a required set of aesthetics to be set. Aesthetic mappings are set with theaes()function. Examples include-
xandy(the position on the x and y axes) -
color(“outside” color, like the line around a bar) -
fill(“inside” color, like the color of the bar itself) -
shape(the type of point, like a dot, square, triangle, etc.) -
linetype(solid, dashed, dotted etc.) -
size(of geoms)
-
2.2 Adding labels and themes
ggplot(data = penguins) +
geom_point(mapping = aes(x = bill_length_mm, y = flipper_length_mm)) +
labs(
x = "Bill length",
y = "Flipper length",
title = "Scatter plot of bill and flipper length",
caption = "R package palmerpenguins"
) +
theme_bw()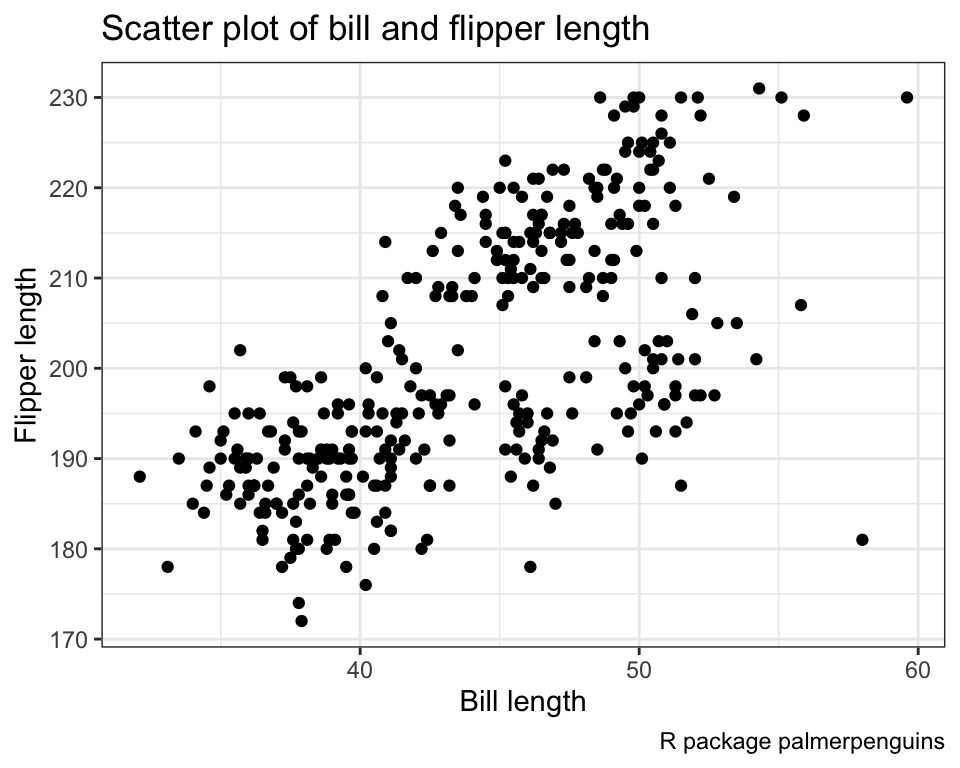
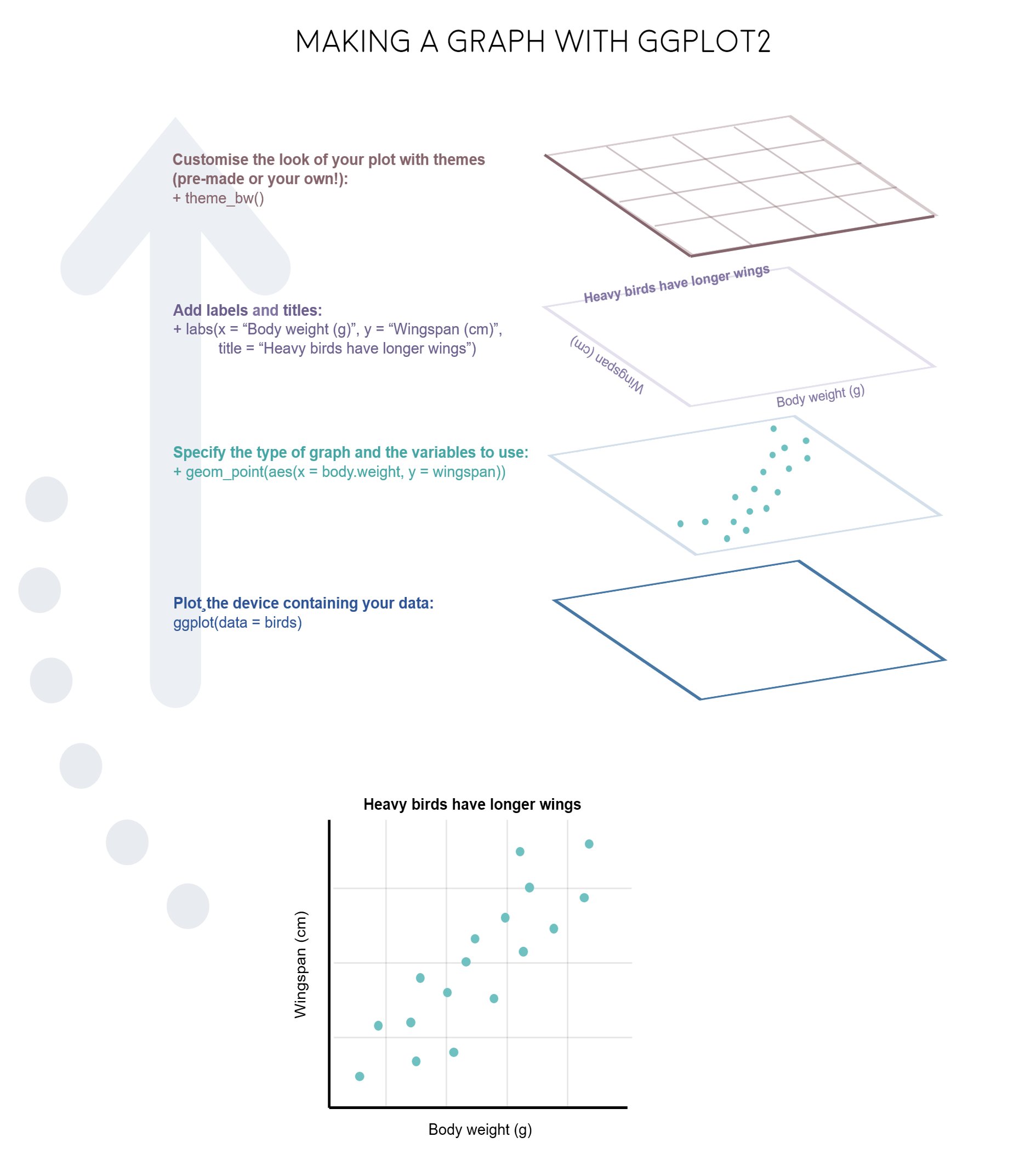
3 Aesthetic Properties in ggplot2
- Variables can be linked to a graph through many aesthetic properties in
ggplot2.
The most commonly used aesthetics are:- colors (
color,fill)
- shapes and lines (
shape,linetype)
- size (
size,linewidth)
- transparency (
alpha)
- group structure (
group)
- position adjustments (
x,y)
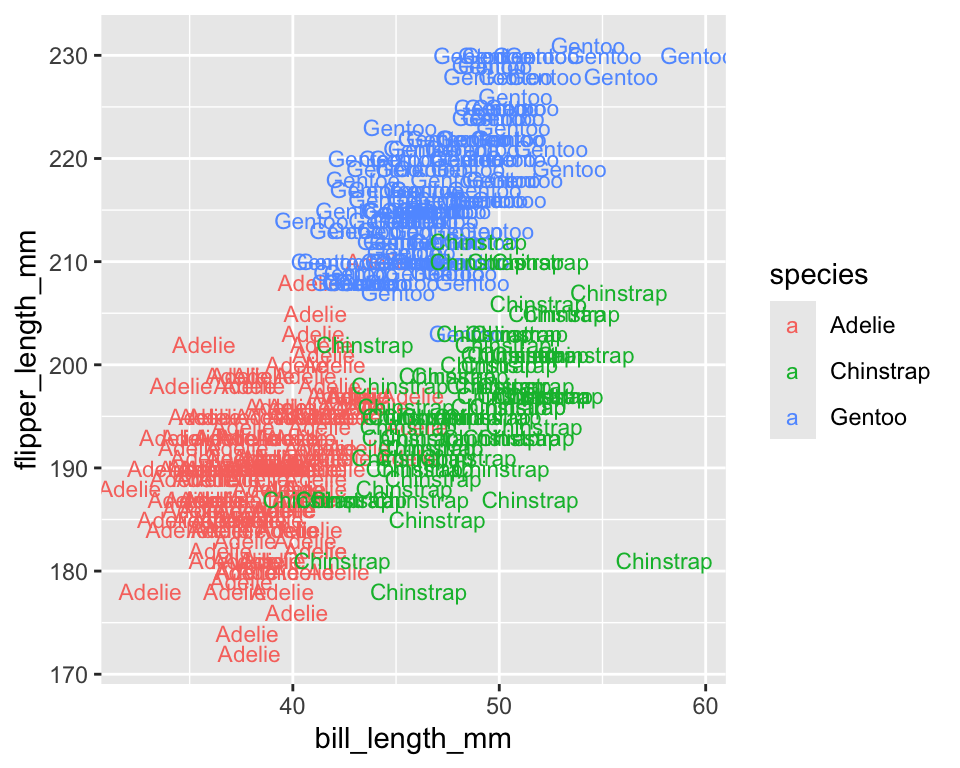
- text aesthetics (
label,family,fontface)
- point-specific (
stroke)
- bar-specific (
width)
- colors (
- Each aesthetic controls a visual aspect of the plot and can be mapped to a variable inside
aes().
-
colorcontrols the outside color of points, lines, and borders - Here different colors represent different levels of
species
ggplot(penguins) +
geom_point(aes(x = bill_length_mm,
y = flipper_length_mm,
color = species))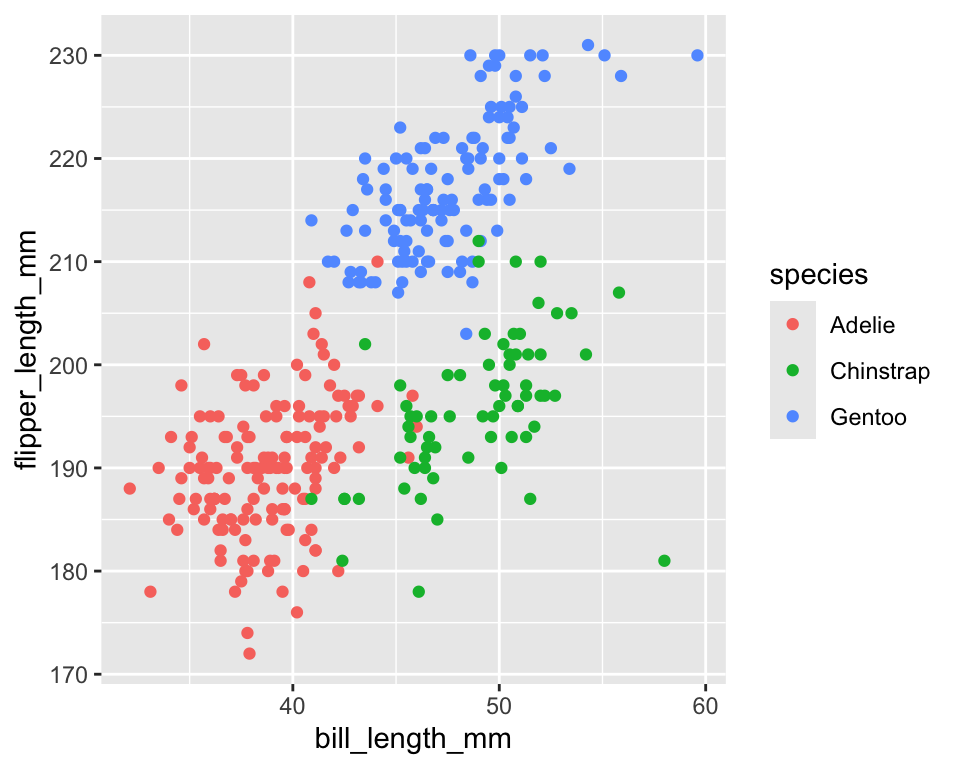
-
fillcontrols the inside color of shapes such as bars and boxes - Works mainly with geoms that have area (bars, boxes, densities)
ggplot(penguins) +
geom_boxplot(aes(x = species, y = body_mass_g, fill = species))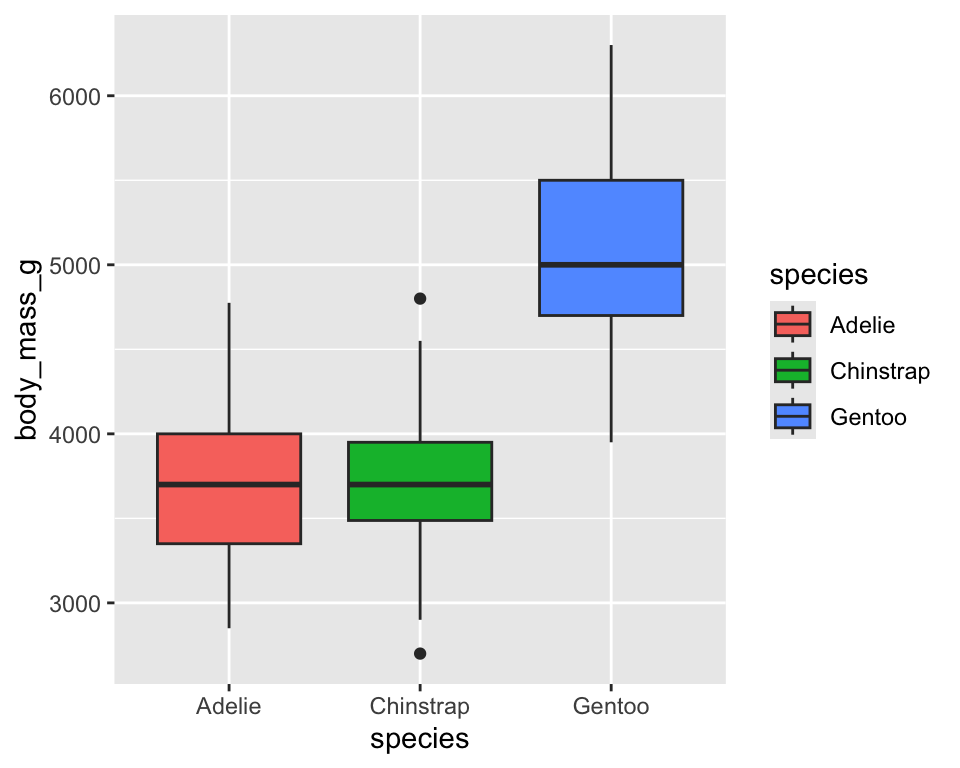
-
shapeassigns different point symbols - ggplot supports up to six distinct shapes for categorical variables
ggplot(penguins) +
geom_point(aes(x = bill_length_mm,
y = flipper_length_mm,
shape = species))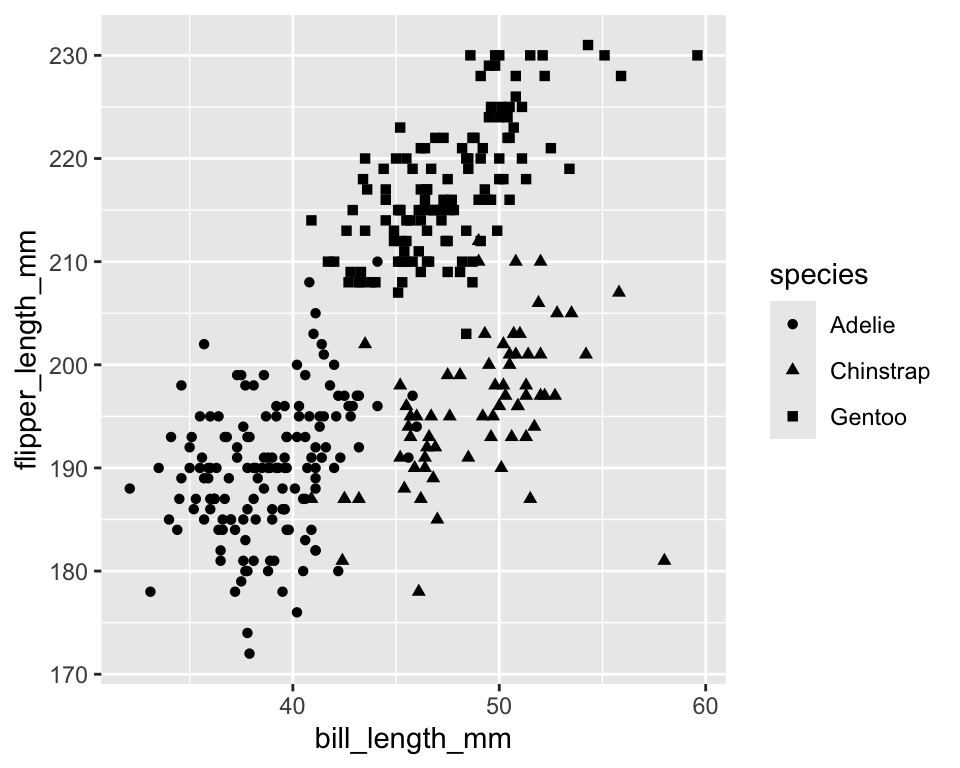
-
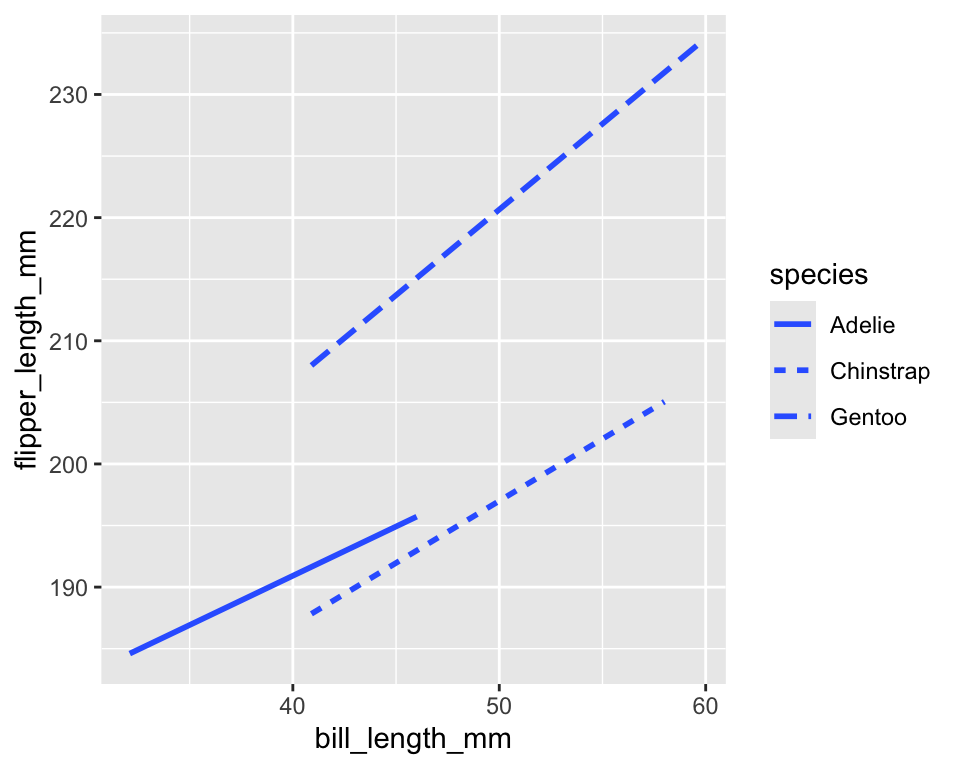
geom_smooth()fits the relationship between two quantitative using a smoothing method -
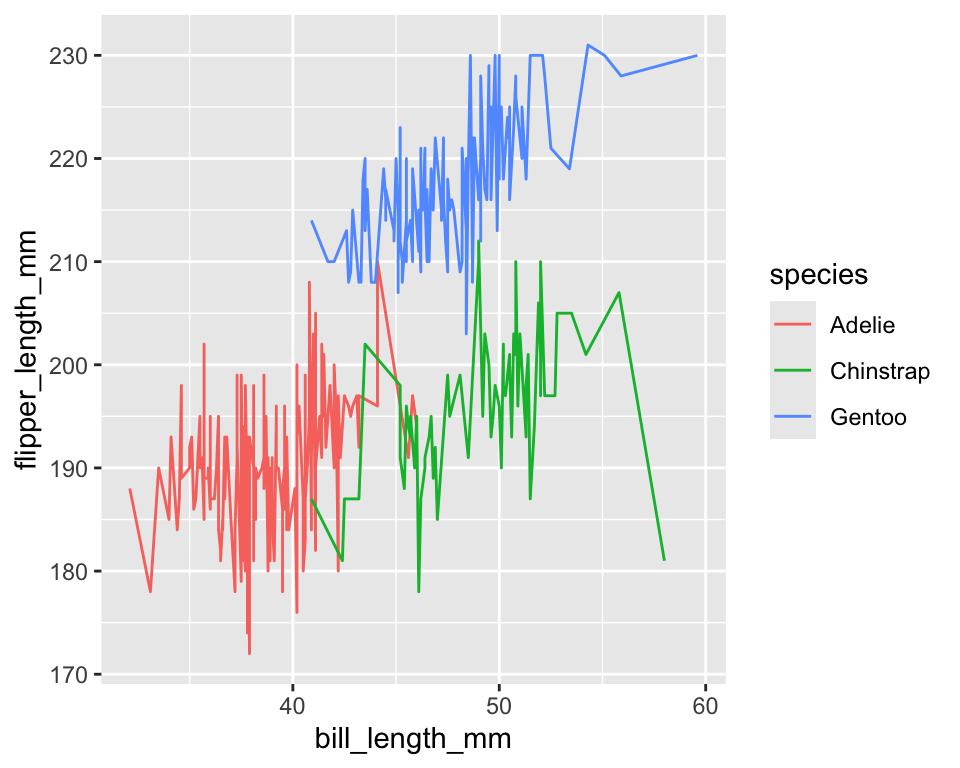
linetypecontrols solid, dashed, dotted patterns - Useful for distinguishing groups in line plots
ggplot(penguins) +
geom_smooth(
aes(x = bill_length_mm, y = flipper_length_mm, linetype = species),
method = "lm",
se = FALSE
)
-
sizecontrols point size
ggplot(penguins) +
geom_point(aes(x = bill_length_mm,
y = flipper_length_mm,
size = body_mass_g))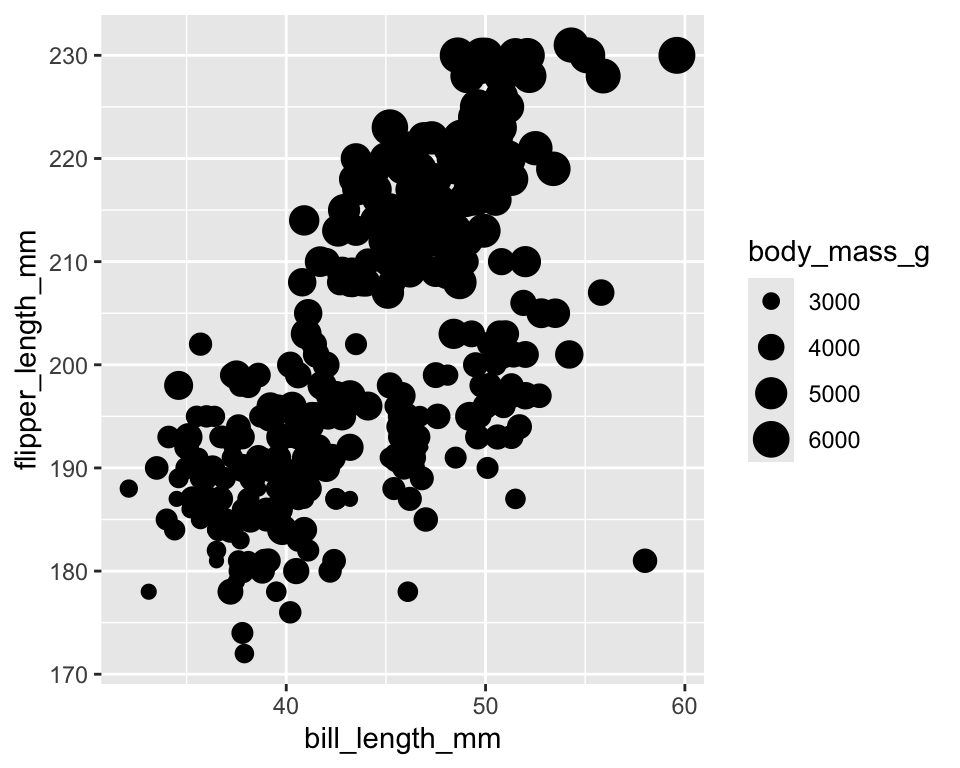
-
alphacontrols transparency (0 = invisible, 1 = opaque) - Useful when points overlap
ggplot(penguins) +
geom_point(aes(x = bill_length_mm,
y = flipper_length_mm,
alpha = species))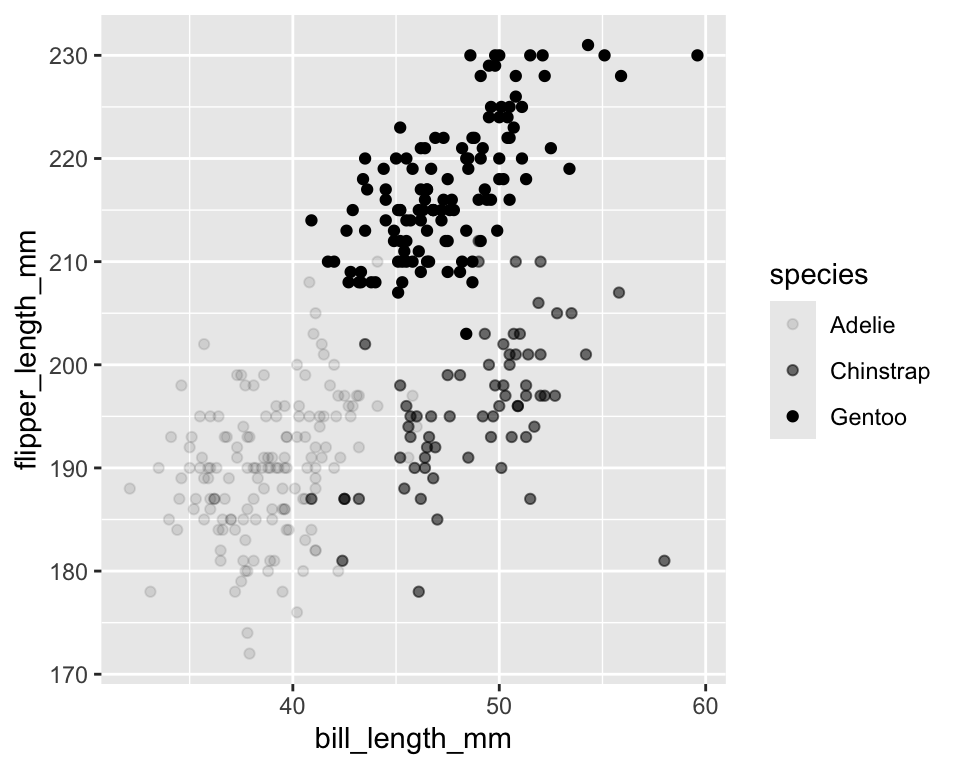
3.1 How to know which aesthetics work with a geom?
- Open the help page:
?geom_point - Use keyboard Tab completion inside
aes()to see suggestions
3.2 Setting vs Mapping aesthetics
- When an aesthetic is inside
aes(), it is mapped to a variable - When it is outside
aes(), it is set to a constant value
# Mapping (data driven)
ggplot(penguins) +
geom_point(aes(bill_length_mm, flipper_length_mm, color = species))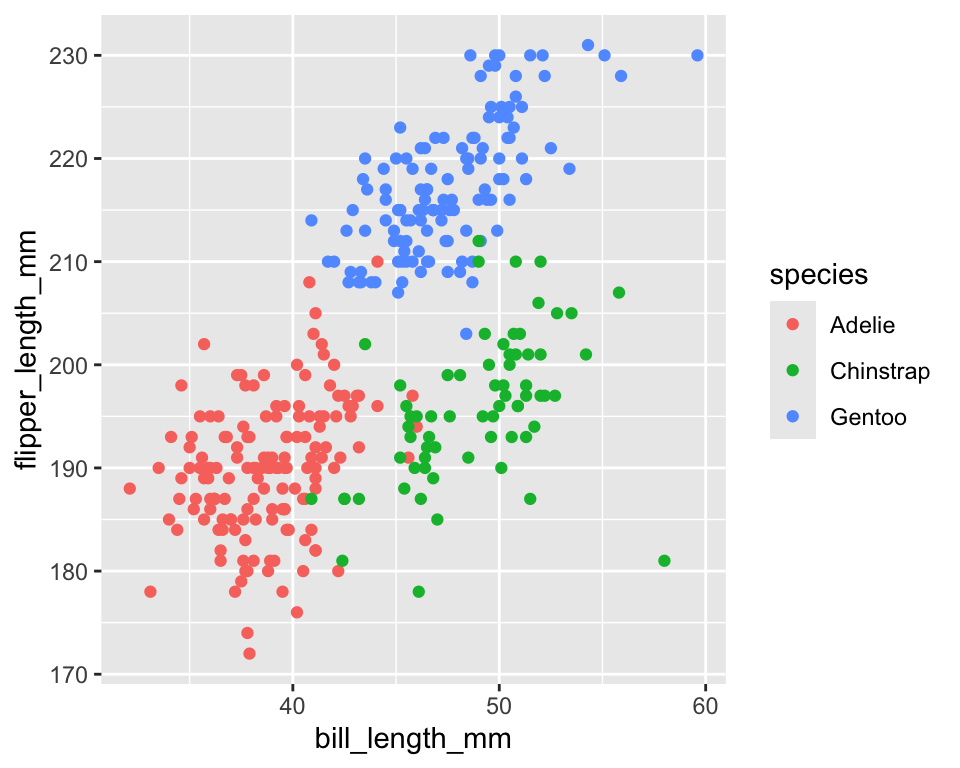
# Setting (fixed appearance)
ggplot(penguins) +
geom_point(aes(bill_length_mm, flipper_length_mm), color = "blue")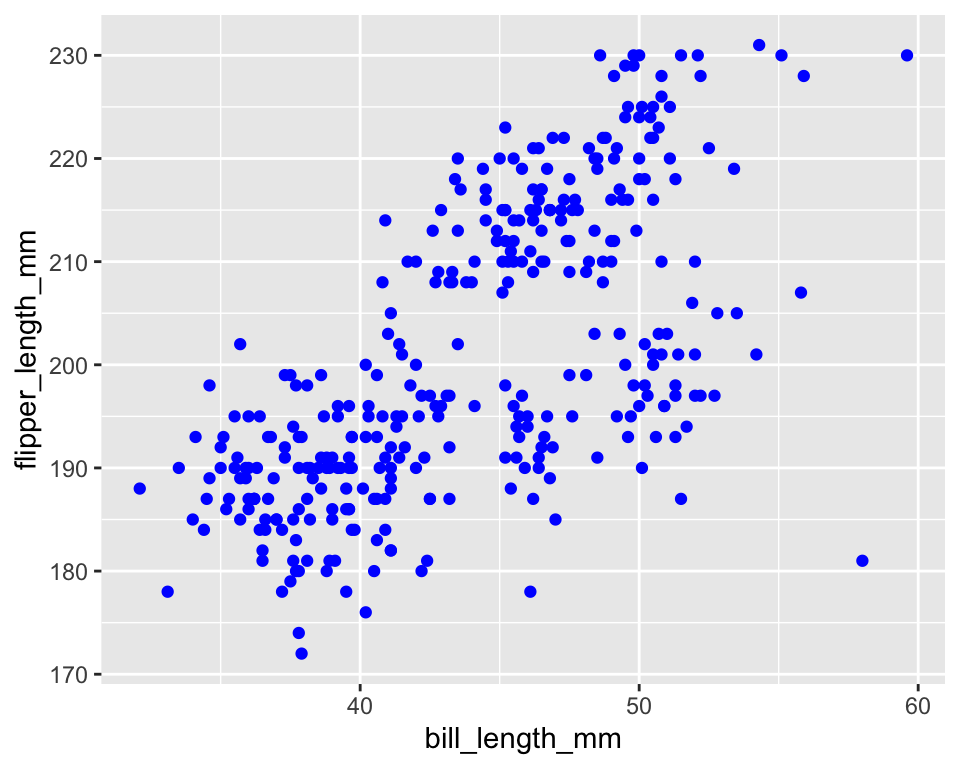
3.3 Exercise 3.2.3
(use diamonds data to answer the followings)
- Create a scatter plot to examine the effect of
priceoncaratand assign different colors to different levels ofcut - Show a fit of a linear model on the scatter plot of
caratandprice - Show different fits of linear models (
priceoncarat) corresponding to different levels ofcuton the scatter plot ofpriceandcarat
4 Histogram
-
geom_histogram()is for histogram - Only
xvalue is needed for itsaes()function
ggplot(penguins) +
geom_histogram(aes(x = body_mass_g))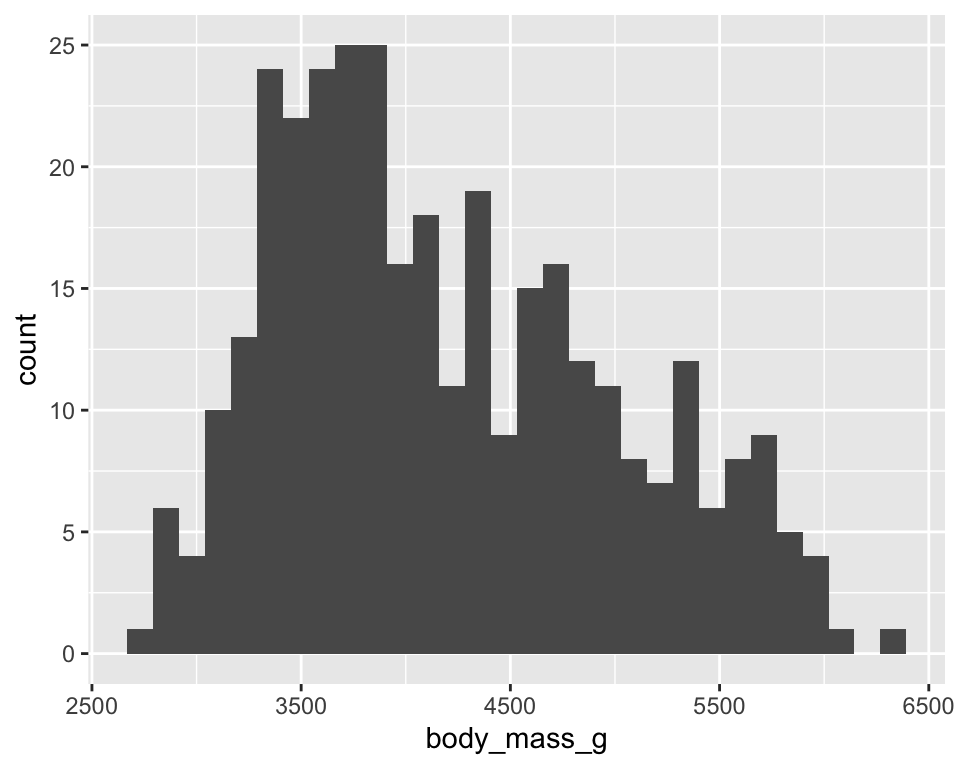
4.1 Histogram and density function
-
geom_density()is used to obtain the density of a variable
ggplot(penguins) +
geom_histogram(aes(x = body_mass_g, y = after_stat(density))) +
geom_density(aes(x = body_mass_g, y = after_stat(density))) 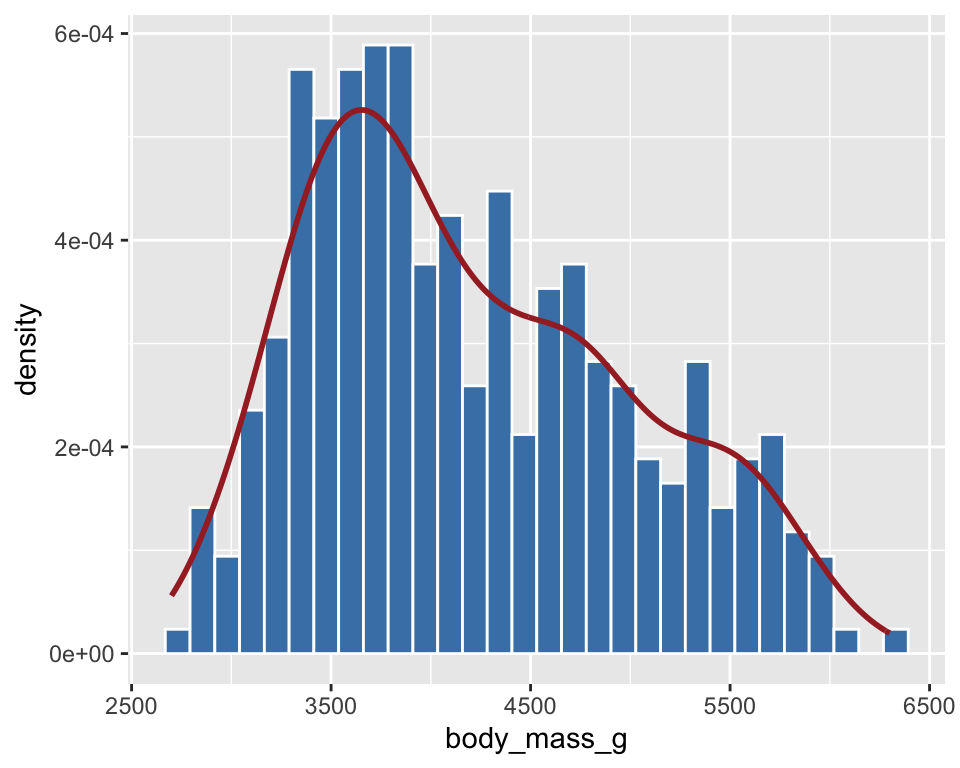
- A common mapping function in
ggplot()for differentgeom_*()
ggplot(penguins, aes(x = body_mass_g, y = after_stat(density))) +
geom_histogram(fill = "steelblue", color = "white") +
geom_density(color = "brown", size = 1) 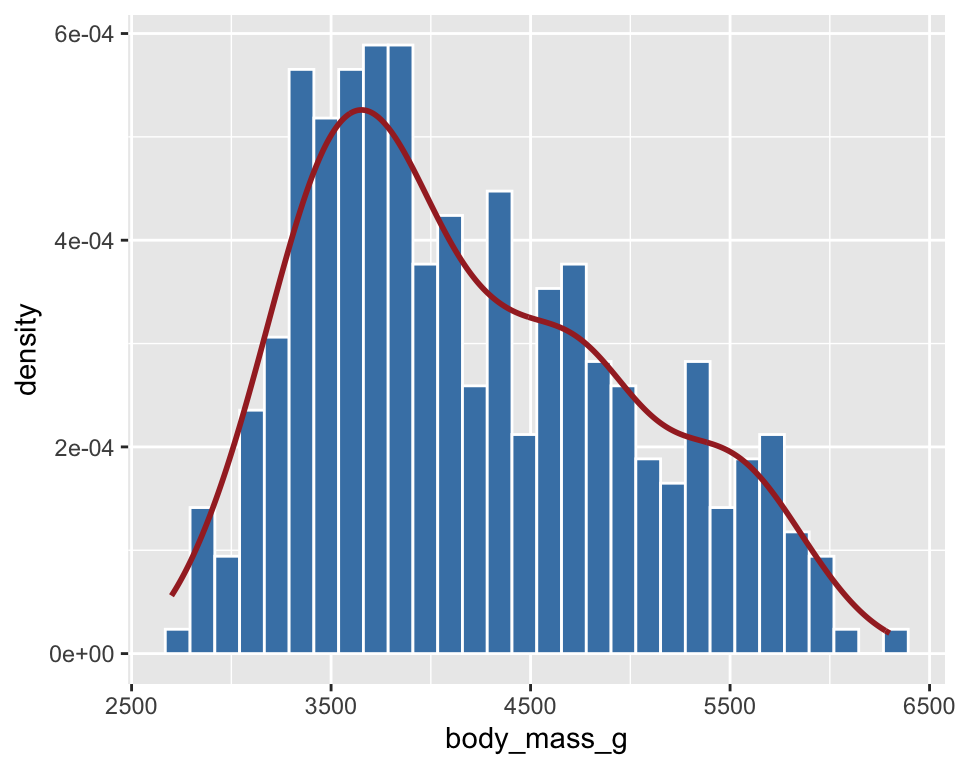
4.2 Exercise 3.2.1
(use diamonds data to answer the followings)
- Create a histogram of
caratand check the effect ofbinson histogram - Add a density line to the plot obtained in Question 1
5 Boxplot
geom_boxplot()
ggplot(penguins) +
geom_boxplot(aes(x = species, y = body_mass_g))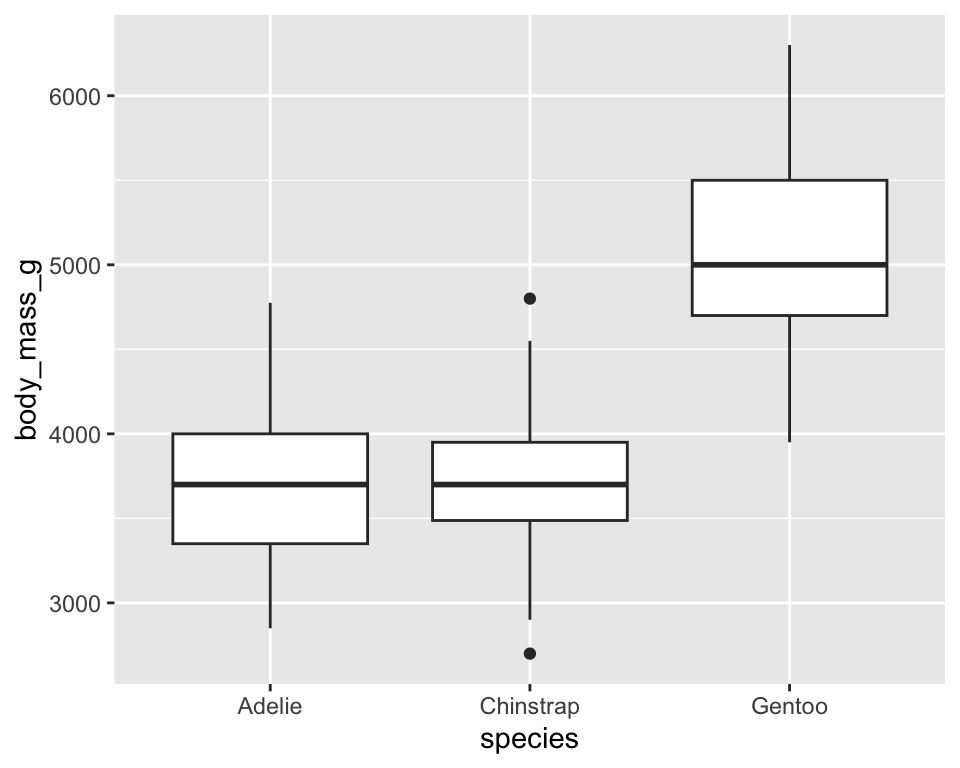
ggplot(penguins) +
geom_boxplot(aes(x = species, y = body_mass_g), fill = "brown") +
coord_flip() 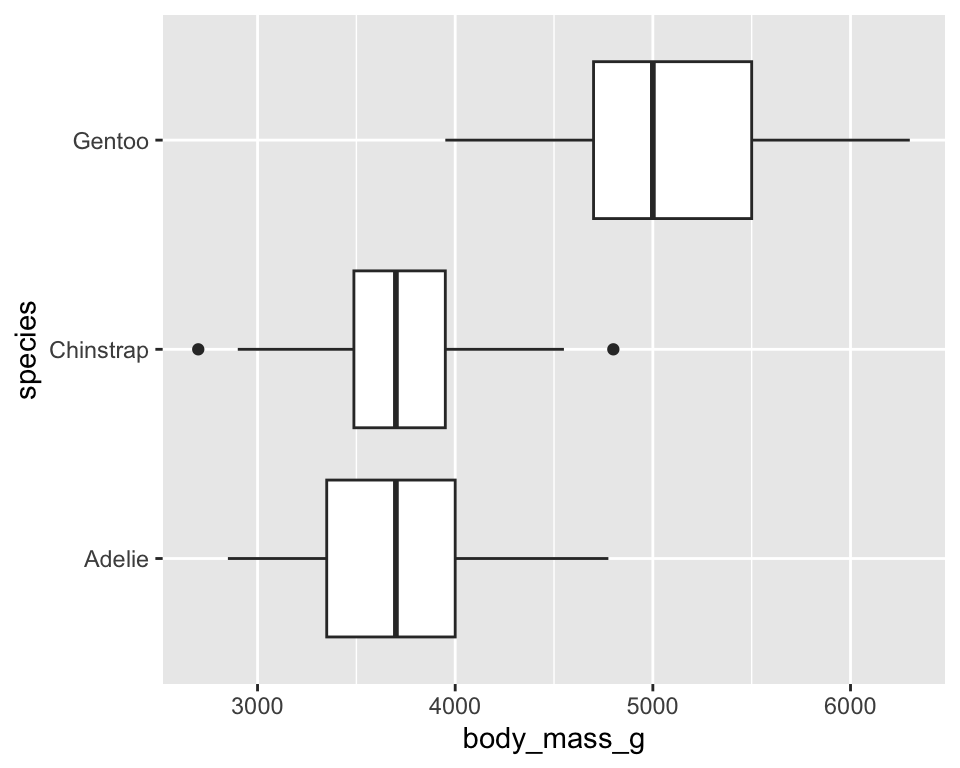
5.1 Boxplot with original data points
ggplot(penguins, aes(x = species, y = body_mass_g)) +
geom_boxplot() +
geom_jitter(width = .2, aes(color = species), size = .75) 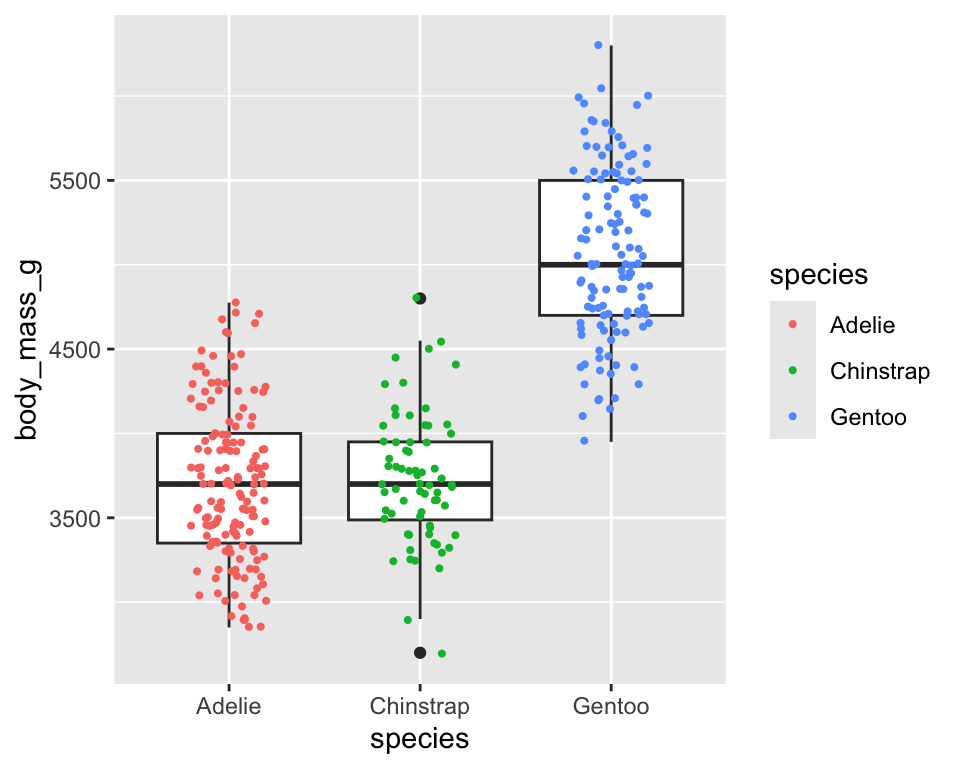
-
geom_jitter()adds a small amount random variation to each point and it is useful to visualize points at different levels
5.2 Exercise 3.2.2
(use diamonds data to answer the followings)
- Create a boxplot of
caratat different levels ofcut - Create a scatter plot to examine the effect of
caratonprice
6 Facets
- Sometimes a plot becomes crowded when too many variables are shown using colors or shapes.
- Facets display additional categorical variables by splitting a plot into multiple smaller panels, one for each group.
- ggplot2 provides two main functions for faceting:
-
facet_wrap()→ splits the plot by one categorical variable -
facet_grid()→ splits the plot by two categorical variables arranged in rows and columns
-
Faceting allows us to compare patterns across groups while keeping the same axes and scales.
ggplot(penguins) +
geom_point(aes(x = flipper_length_mm, y = bill_length_mm)) +
facet_wrap(~species) 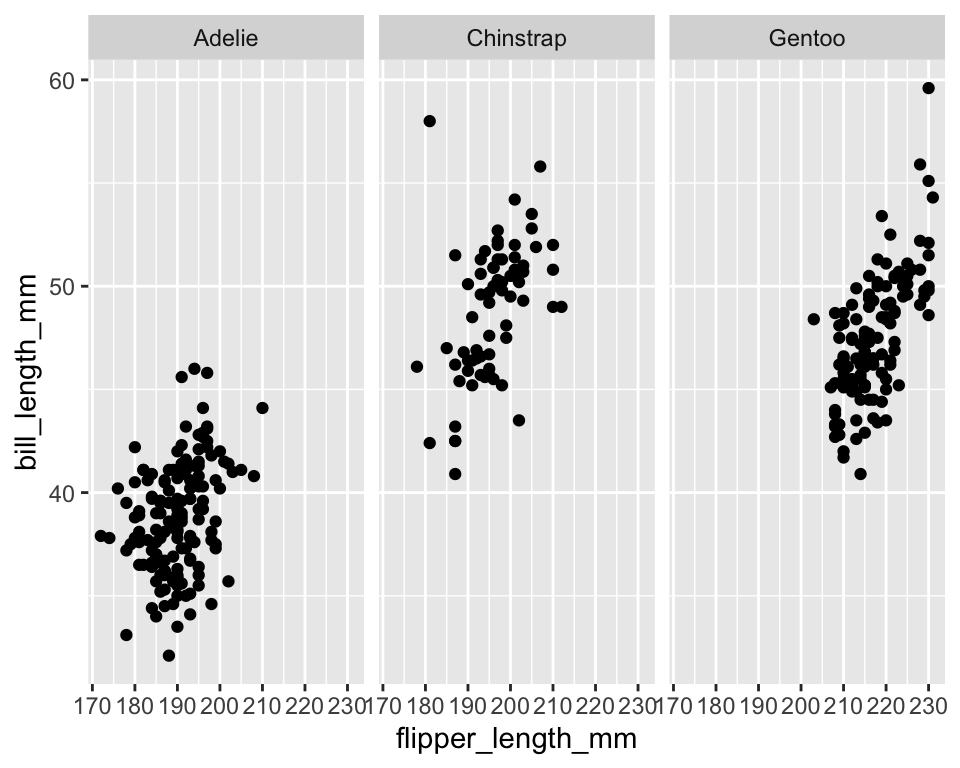
ggplot(penguins,
aes(x = flipper_length_mm, y = bill_length_mm, color = species)) +
geom_point() +
geom_smooth(method = "lm", se = FALSE, color = "black") +
facet_wrap(~species) 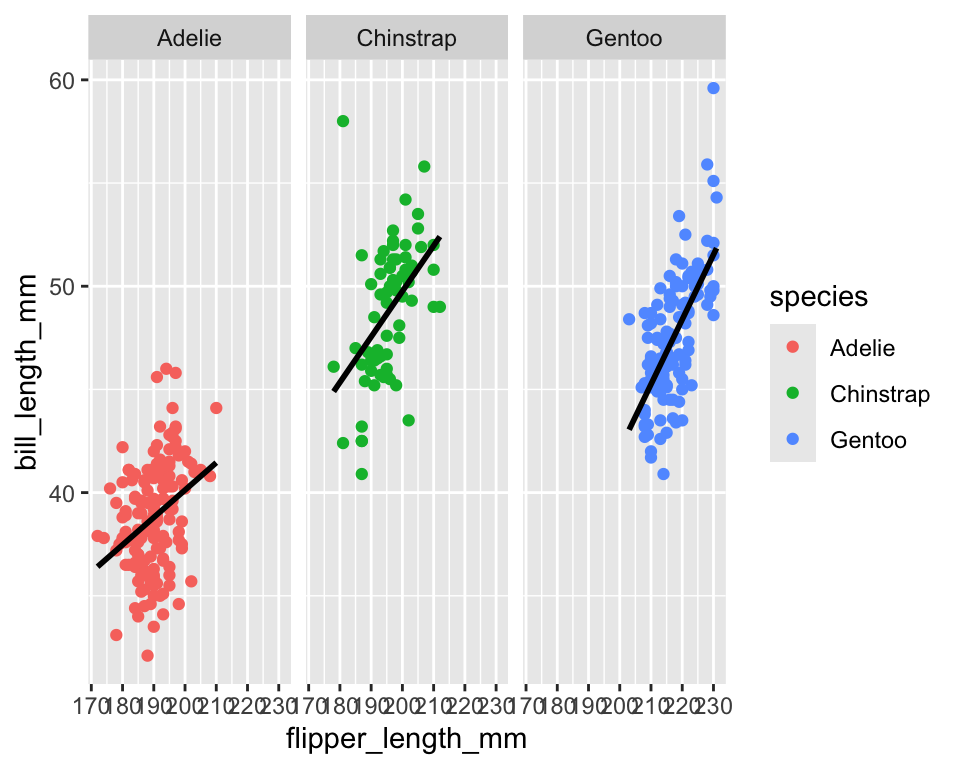
ggplot(data = penguins[!is.na(penguins$sex), ]) +
geom_point(aes(x = flipper_length_mm, y = bill_length_mm)) +
facet_wrap(~ sex + species) 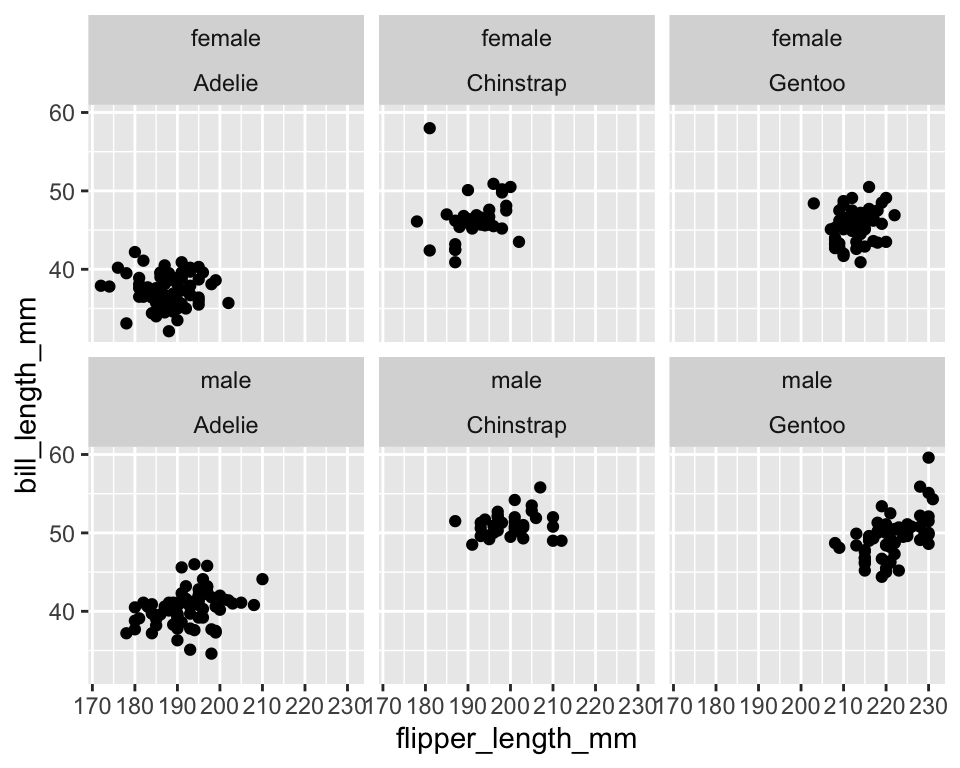
ggplot(data = penguins[!is.na(penguins$sex), ]) +
geom_point(aes(x = flipper_length_mm, y = bill_length_mm)) +
facet_grid(sex ~ species) 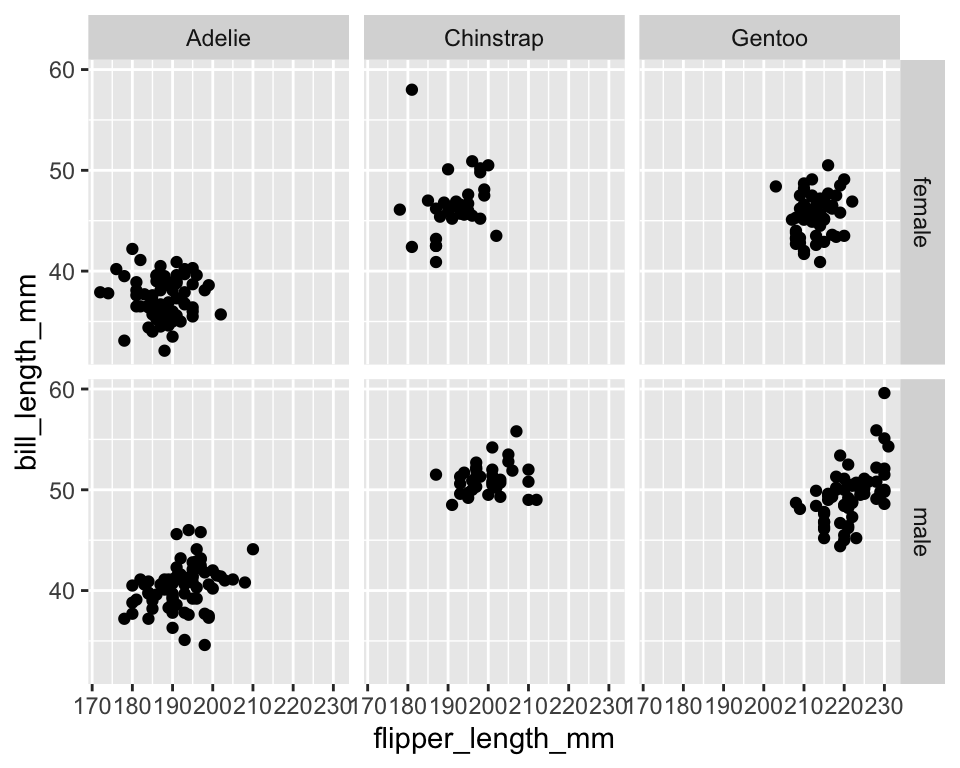
ggplot(penguins) +
geom_histogram(aes(x = body_mass_g),
color = "brown",
fill = "yellow") +
facet_wrap(~ species, ncol = 1)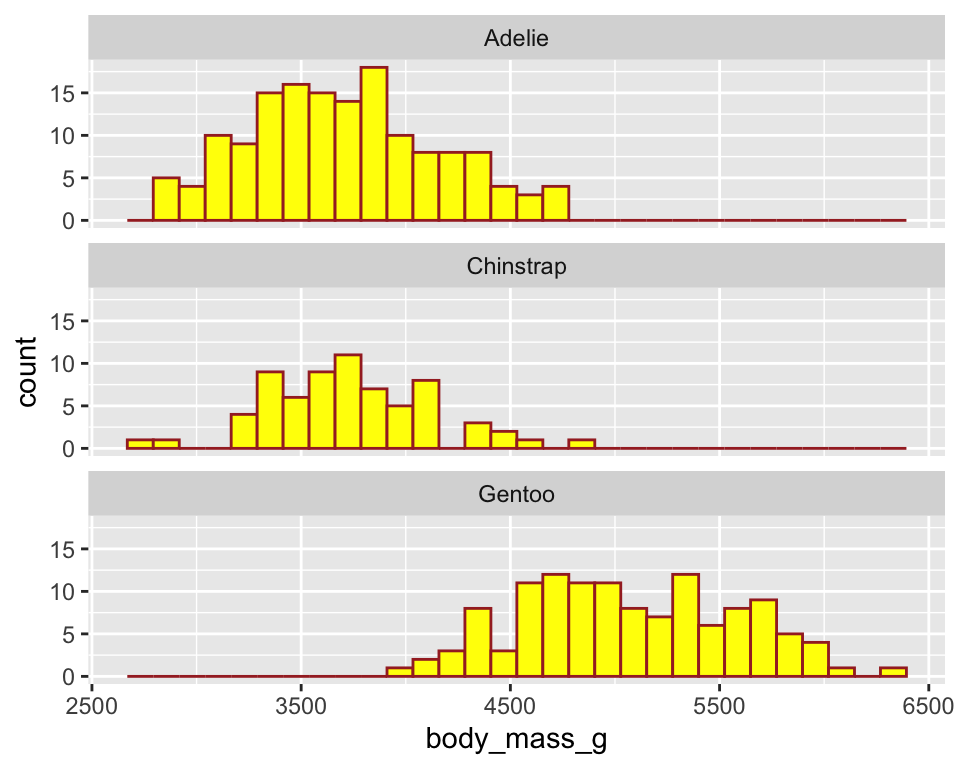
ggplot(penguins) +
geom_histogram(aes(x = body_mass_g),
color = "brown",
fill = "yellow") +
facet_wrap(~ species, ncol = 1) +
theme_minimal() 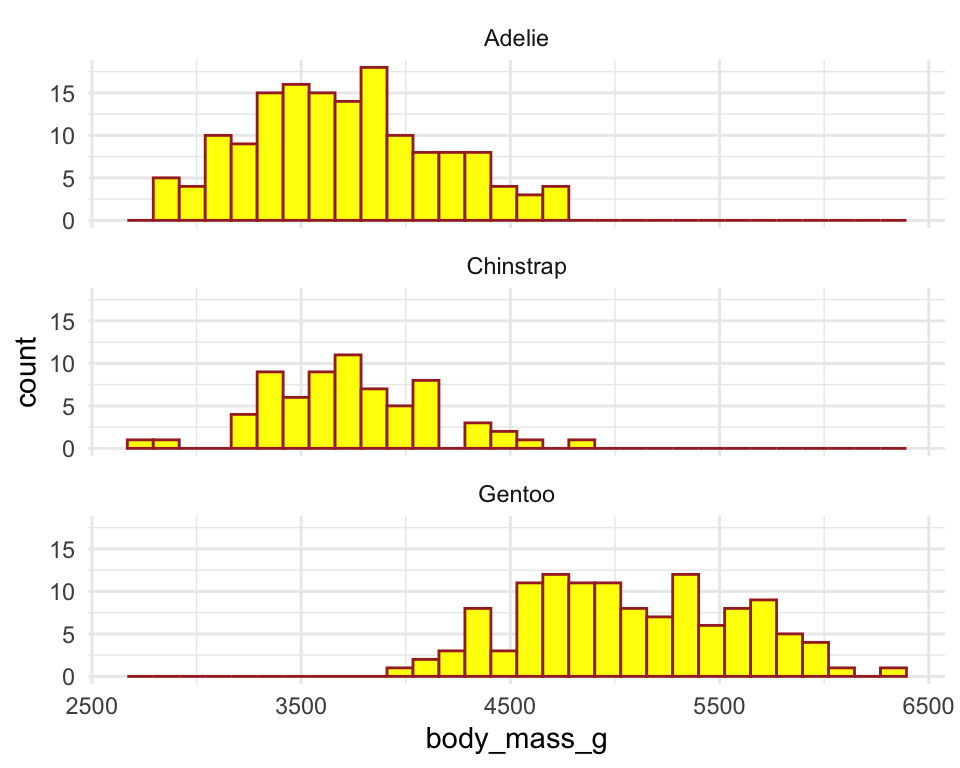
ggplot(penguins) +
geom_histogram(aes(x = body_mass_g),
color = "brown",
fill = "yellow") +
facet_wrap(~ species, ncol = 1) +
theme_minimal() +
theme(panel.grid = element_blank())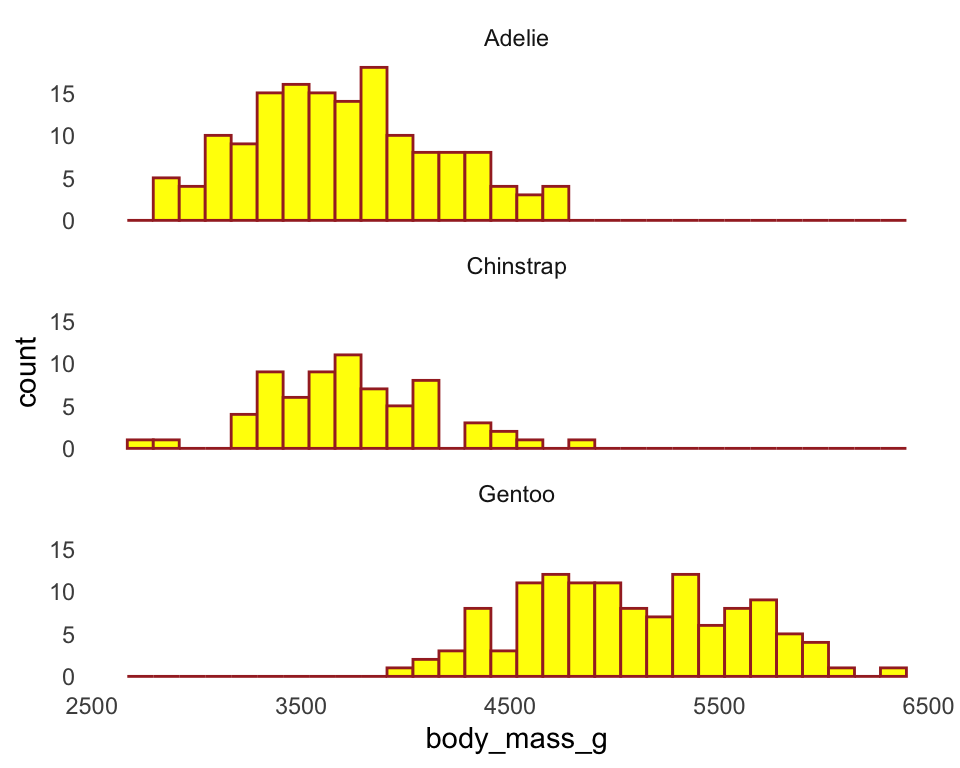
6.1 Exercise 3.2.4
(use diamonds data to answer the followings)
- Create histogram of
xat different levels ofcut
7 Density plot
Distribution of penguins’ body mass
ggplot(penguins) +
geom_density(aes(x = body_mass_g, fill = species), alpha = .5) 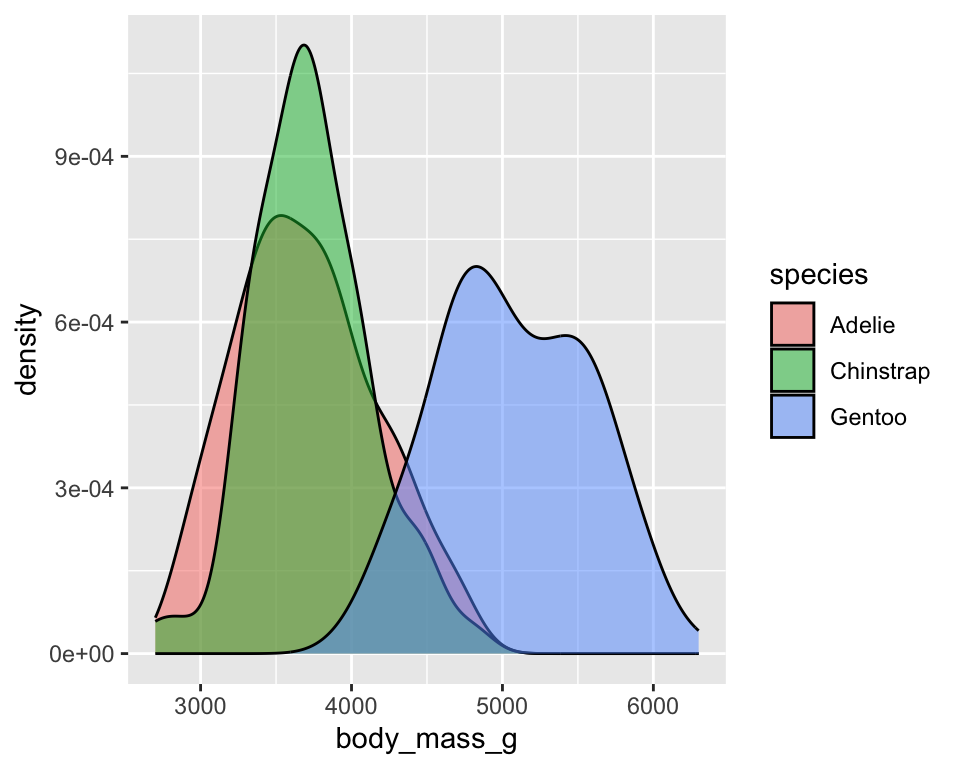
Distribution of penguins’ body mass
ggplot(penguins) +
geom_density(aes(x = body_mass_g, fill = species), alpha = .5) +
theme(legend.position = "top", legend.title = element_blank())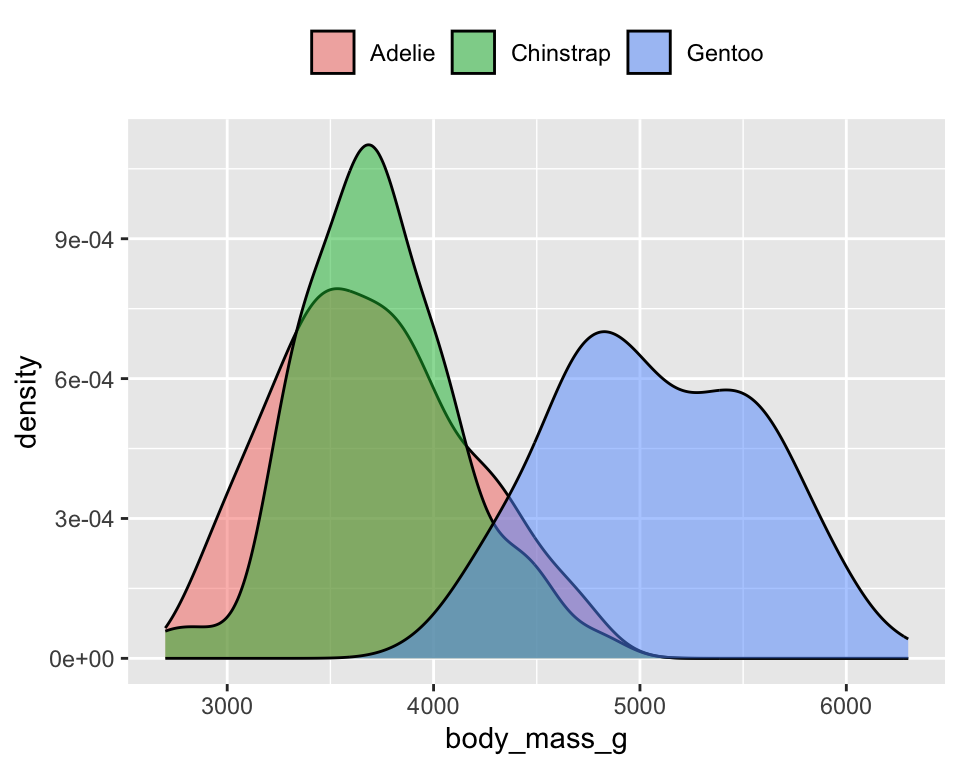
Distribution of penguins’ body mass
ggplot(penguins) +
geom_density(aes(x = body_mass_g, fill = species), alpha = .5) +
theme_minimal(base_size = 7) +
theme(legend.position = "top", legend.title = element_blank())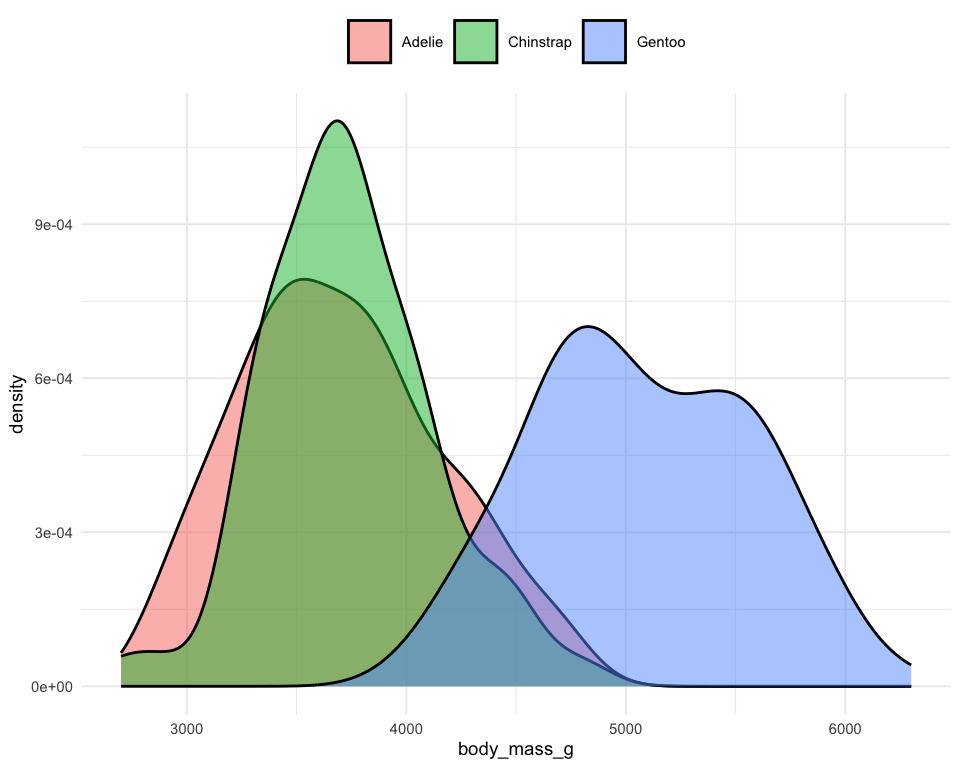
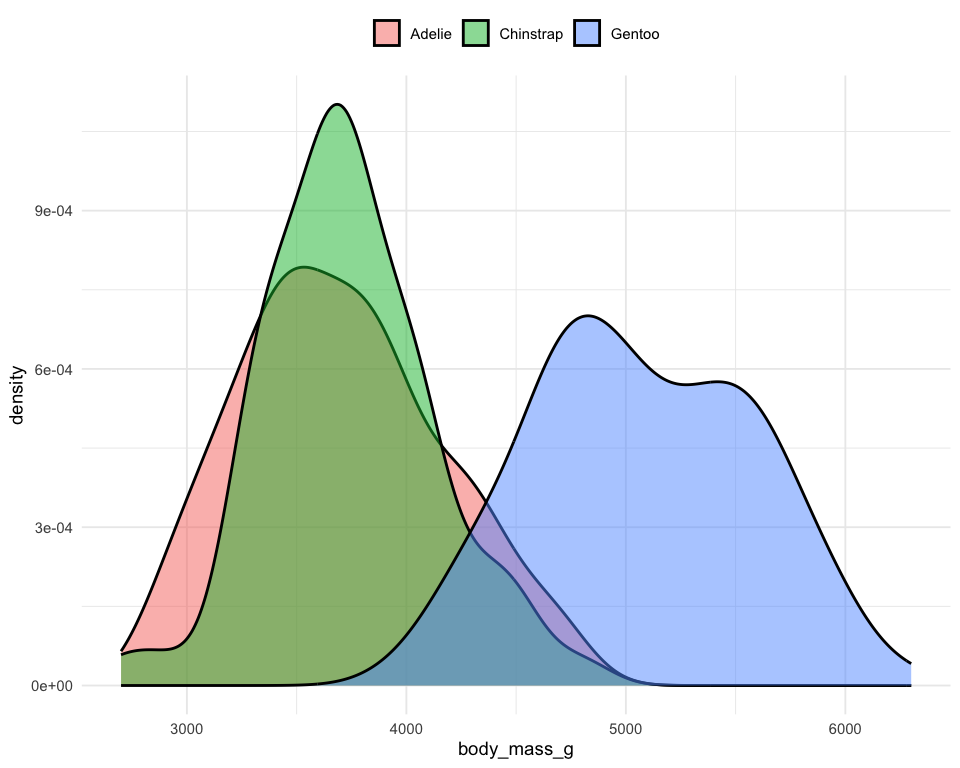
Distribution of penguins’ body mass
ggplot(penguins) +
geom_density(aes(x = body_mass_g, fill = species), alpha = .5) +
theme_minimal(base_size = 7) +
theme(
legend.position = "top",
legend.key.size = unit(.75, "lines"),
legend.title = element_blank()
)
8 Bar Chart
8.1 Frequency bar chart
Frequency distribution of penguin species
8.2 Value bar chart (mean)
Mean Body Mass by Species
penguins |>
group_by(species) |>
summarise(mean_mass = mean(body_mass_g, na.rm=T)) |>
ggplot(aes(x = species, y = mean_mass)) +
geom_col(fill = "steelblue")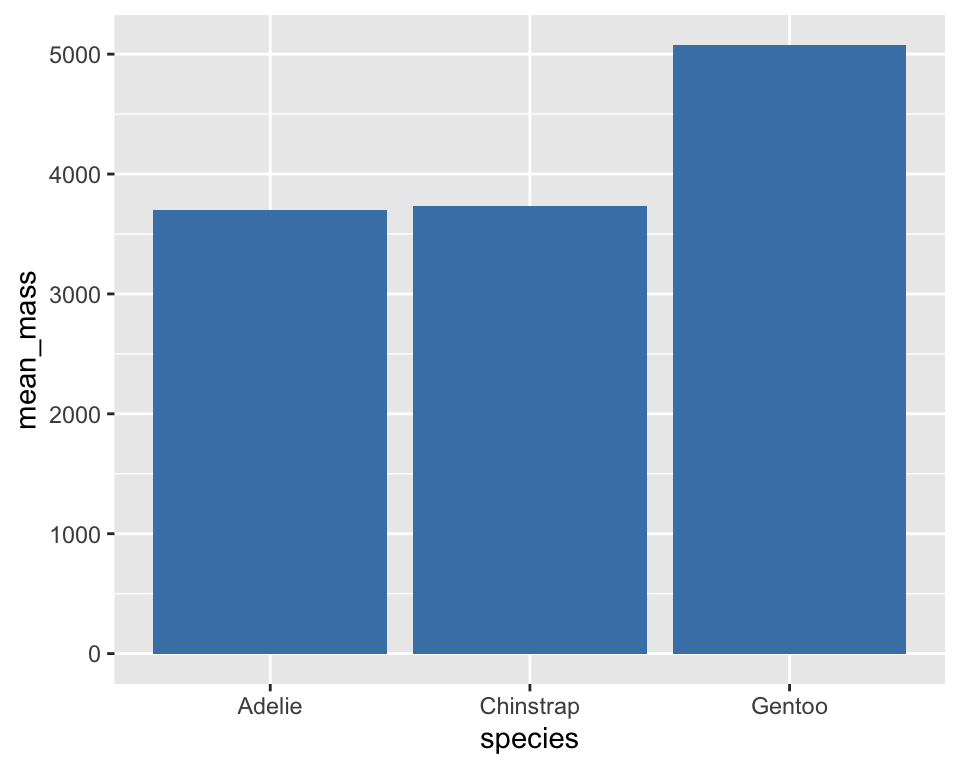
-
geom_col()creates a bar chart where bar heights come from a variable, whilegeom_bar()creates a bar chart based on frequencies.
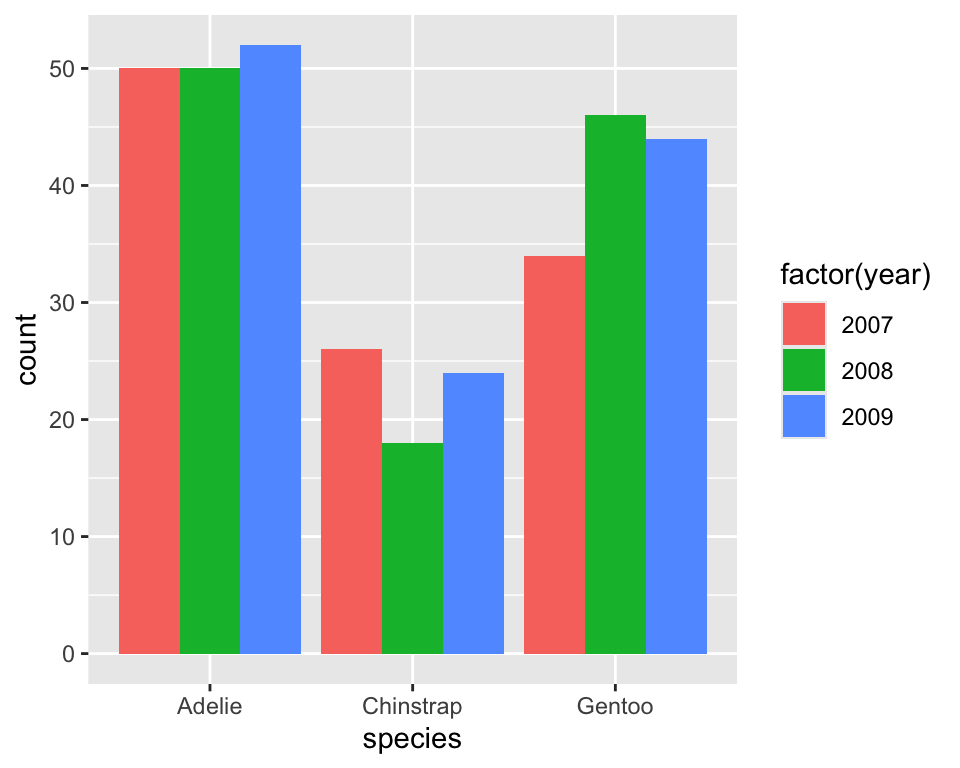
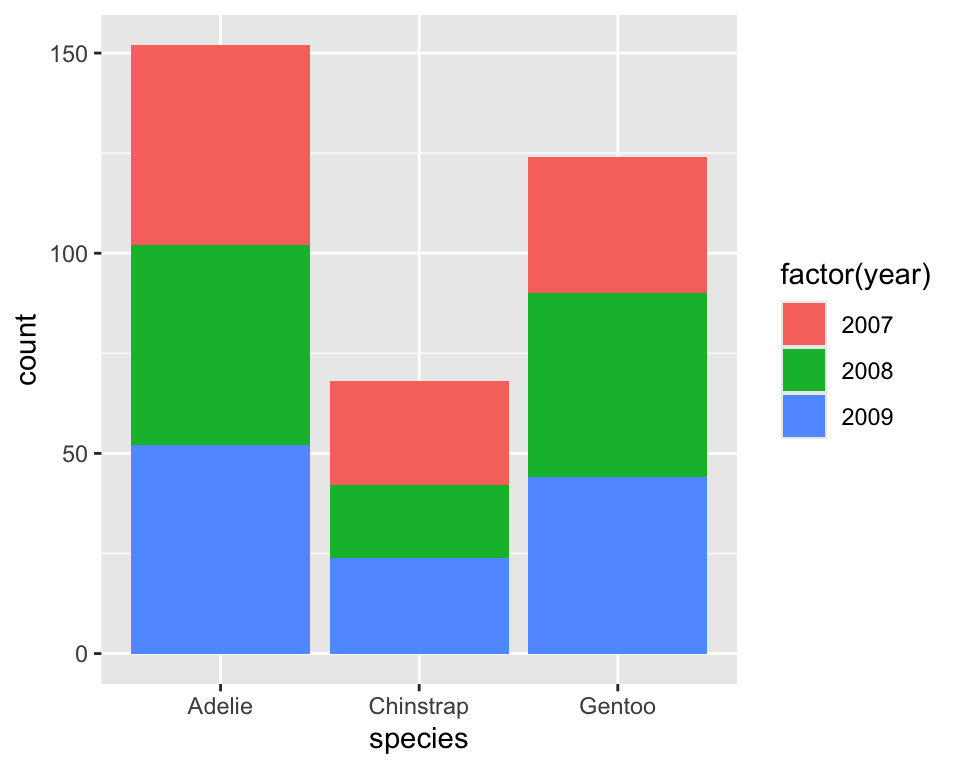
8.3 Frequency barchart with two variables
Distribution of species by year
8.4 Exercise 3.2.5
(use diamonds data to answer the followings)
- Create a barplot of
cut - Create a barplot of
color - Create a barplot of
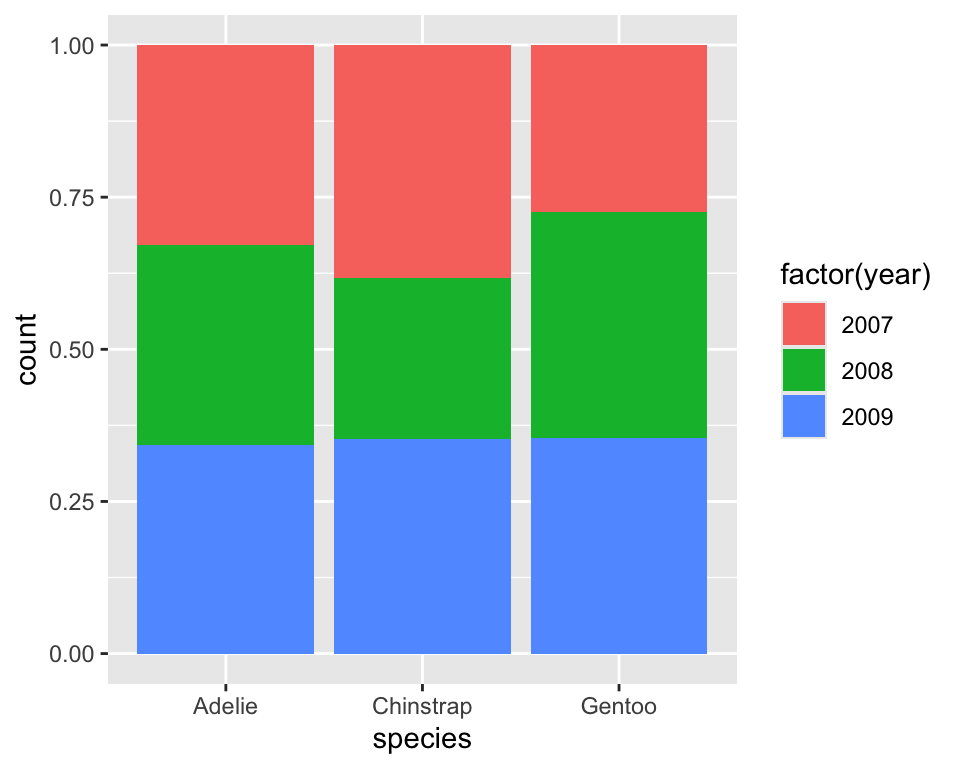
cutwith showing the distribution ofcolorat different levels ofcut - Check the use three different value of the argument
positionwhen creating a barplot withcutandcolor
9 Homework
- Use the package
gapminderto get an access to the datagapminder -
gapminderhas 6 variables and 1704 observations, where the variables are:
#> [1] "country" "continent" "year" "lifeExp" "pop" "gdpPercap"- Create a scatter plot to examine how
gdpPercapaffectslifeExp - Change the scale of x-axis to log base 10
- Add a color layer corresponding to
continentto the previous graph
- Create a scatter plot of
gdpPercapversuslifeExpfor different continents in different plotting regions - Add smooth lines to describe relationship between
gdpPercapandlifeExpfor different continents separately - Draw a boxplot of
lifeExpto compare distribution life expectancy for different continents - Draw a histogram of
lifeExpand check it shapes for different bin size - Draw density plots of
lifeExpfor different continents in a single plot
- Make a scatter plot of
lifeExpon the y-axis againstyearon thex - Fit a straight line to estimate mean life expectancy for a year for different countries
- Split the plot for different continents
- Add a continent-specific mean line to the plot
10 Statistical layers geom_*() vs stat_*()
- In ggplot2, every geom is linked to a statistic.
- Each geom has a default stat, and each stat has a default geom.
10.1 Example: smoothing
# The following codes produce the same result.
ggplot(penguins, aes(bill_length_mm, flipper_length_mm)) +
geom_smooth(stat = "smooth")ggplot(penguins, aes(bill_length_mm, flipper_length_mm)) +
stat_smooth(geom = "smooth")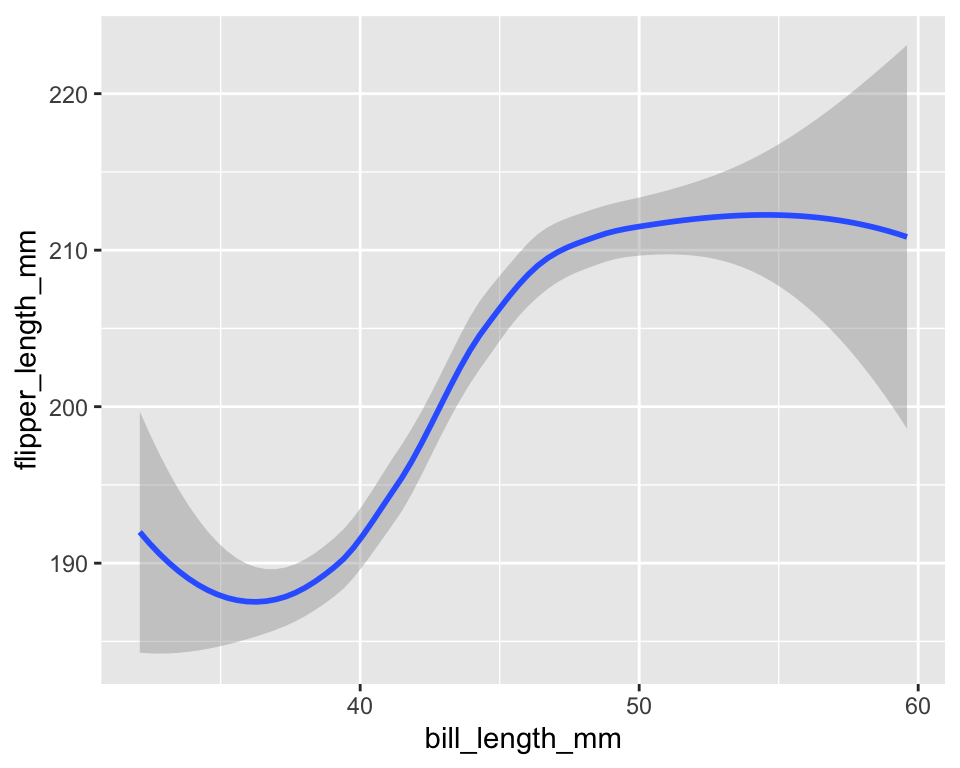
10.2 Example: identity stat for points
# The following codes produce the same result.
ggplot(penguins, aes(bill_length_mm, flipper_length_mm)) +
geom_point(stat = "identity")ggplot(penguins, aes(bill_length_mm, flipper_length_mm)) +
stat_identity(geom = "point")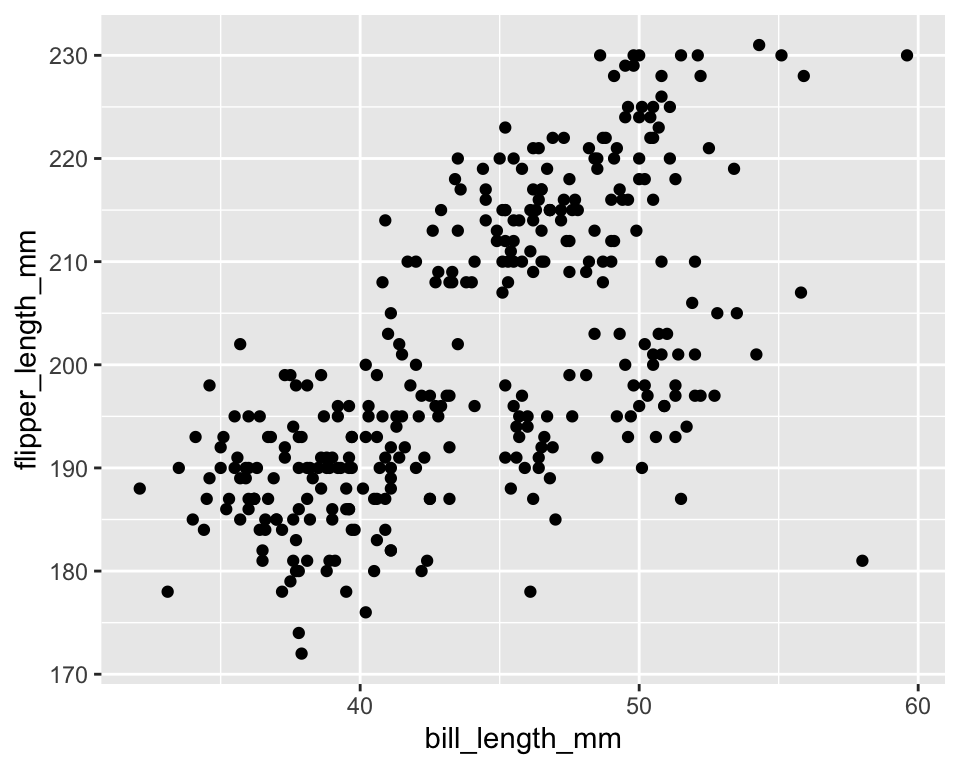
10.3 Example: counting for bar charts
ggplot(penguins, aes(species)) +
stat_count(geom = "bar")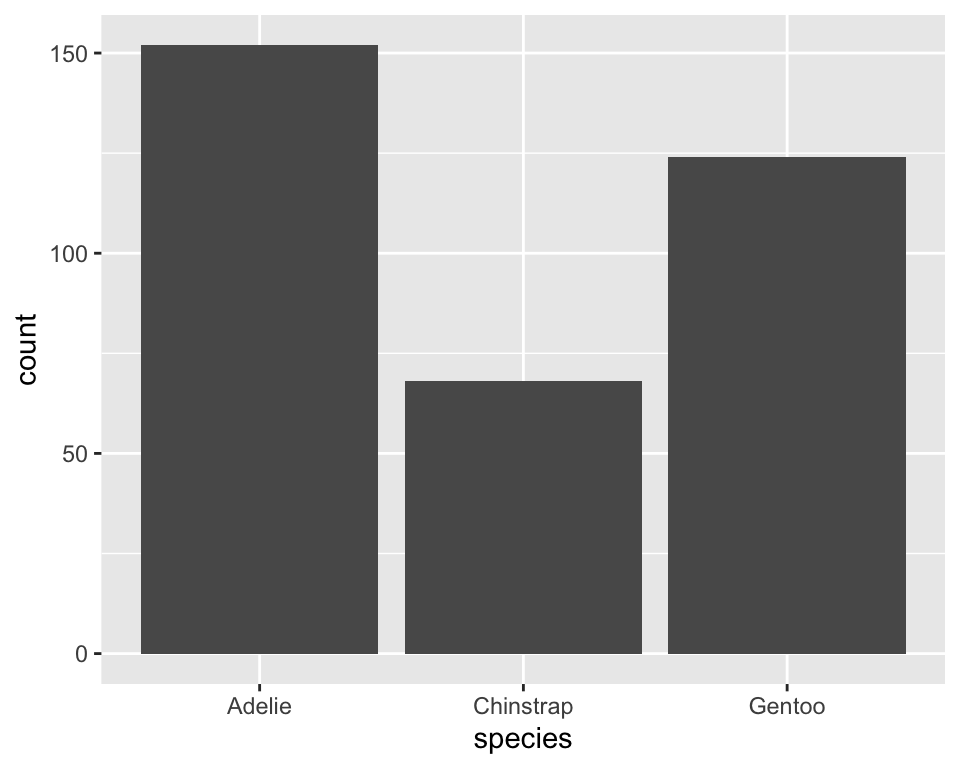
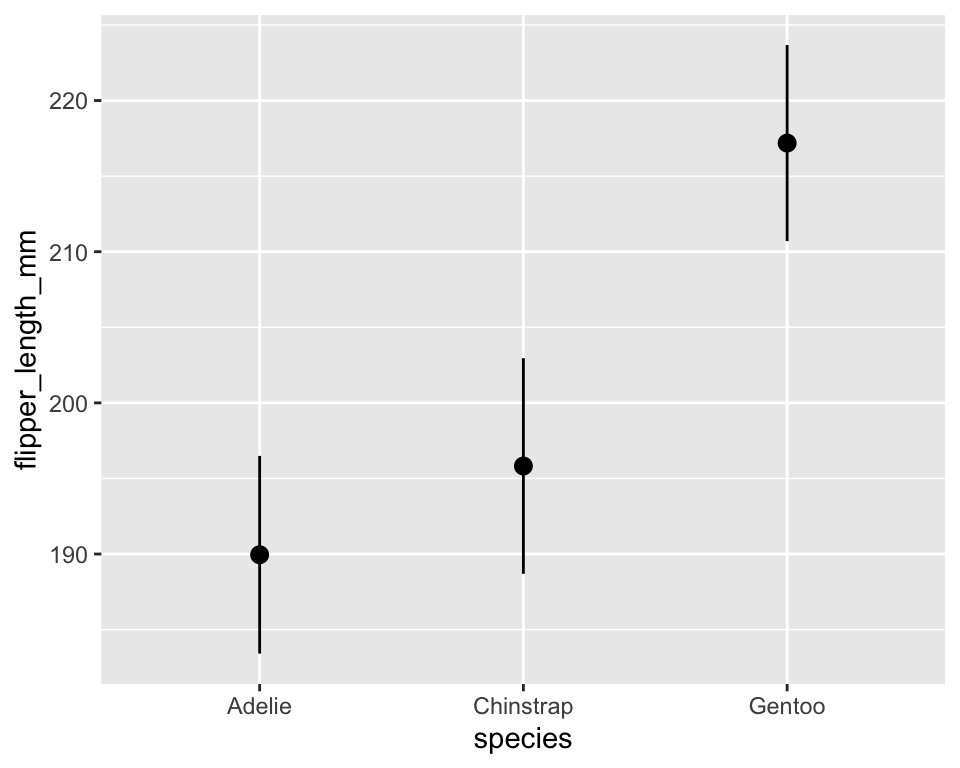
10.4 Statistical summaries
-
stat_summary()allows us to display custom summaries instead of raw data.
# The following codes produce the same result.
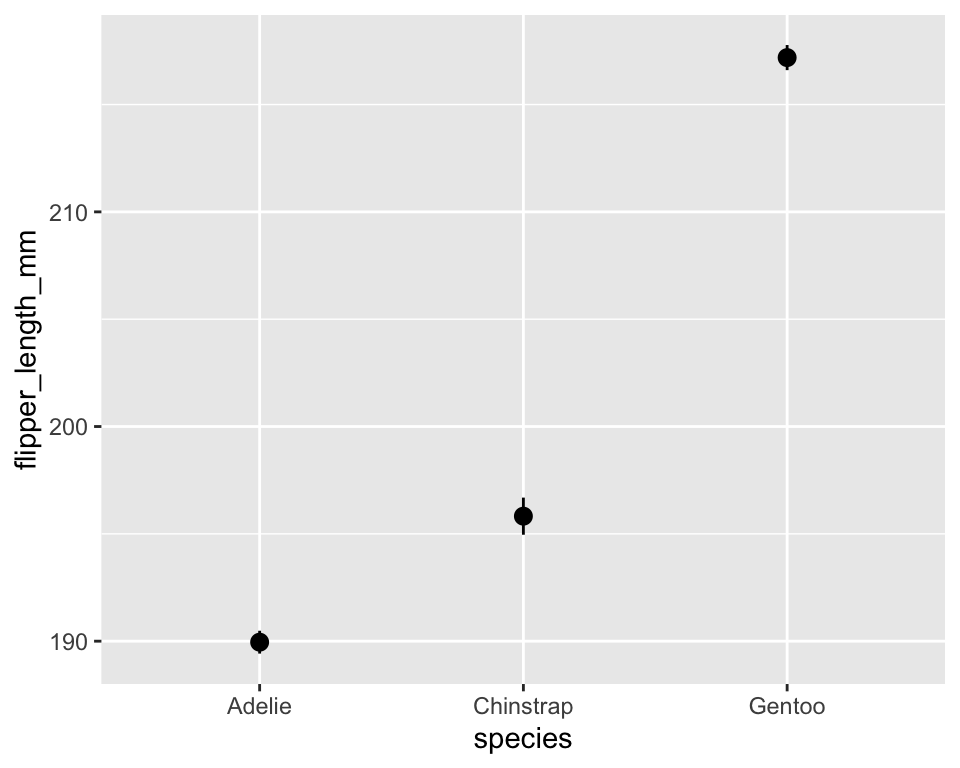
ggplot(penguins, aes(species, flipper_length_mm)) +
stat_summary()ggplot(penguins, aes(species, flipper_length_mm)) +
stat_summary(fun.data = mean_se, geom = "pointrange")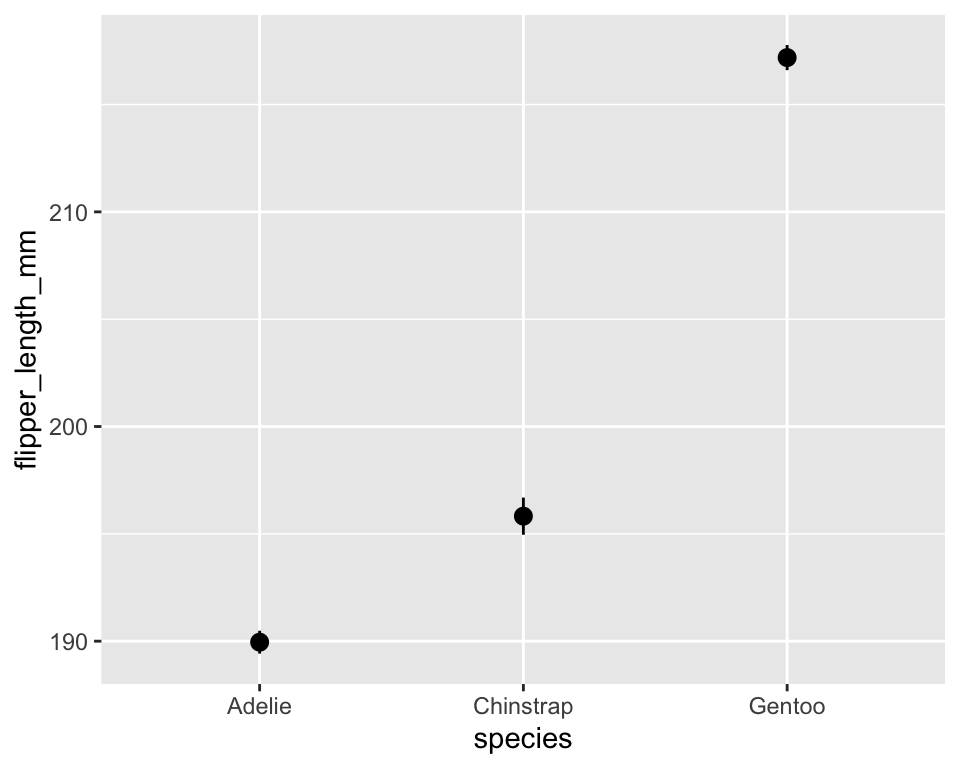
Adding summaries to existing geoms
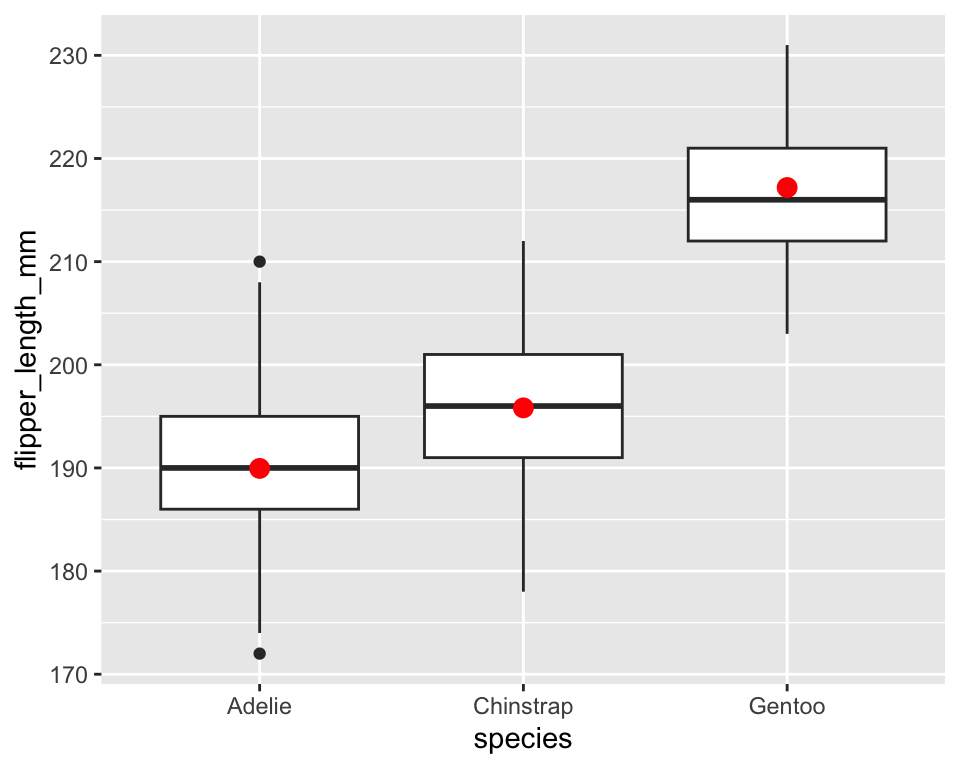
ggplot(penguins, aes(species, flipper_length_mm)) +
geom_boxplot() +
stat_summary(
fun = mean,
geom = "point",
color = "red",
size = 3
)
Mean Body Mass by Species
ggplot(penguins, aes(x = species, y = body_mass_g)) +
stat_summary(fun = mean, geom = "col", fill = "steelblue")