library(purrr) # library(tidyverse) would also work17 {purrr}
Note
This lecture used materials from the iteration chapter in R for data science.
The map functions
The pattern of looping over a vector, doing something to each element and saving the results is so common that the purrr package provides a family of functions to do it for you. There is one function for each type of output:
map()makes a list.map_lgl()makes a logical vector.map_int()makes an integer vector.map_dbl()makes a double vector.map_chr()makes a character vector.
Each function takes a vector as input, applies a function to each piece, and then returns a new vector that’s the same length (and has the same names) as the input. The type of the vector is determined by the suffix to the map function.
- Imagine we have this simple tibble:
df <- tibble(
a = rnorm(10),
b = rnorm(10),
c = rnorm(10),
d = rnorm(10)
)- We want to compute the mean, median, and SD of each column
map_dbl(df, mean)## a b c d
## -0.50627203 -0.42086929 -0.40903136 0.08975649map_dbl(df, median)## a b c d
## -0.2855340 -0.5648310 -0.3025734 -0.2135869map_dbl(df, sd)## a b c d
## 0.6171634 0.7422437 1.0691040 0.9295846- Compared to using a
forloop, focus is on the operation being performed (i.e.mean(),median(),sd()), not the bookkeeping required to loop over every element and store the output. This is even more apparent if we use the pipe:
df |> map_dbl(mean)## a b c d
## -0.50627203 -0.42086929 -0.40903136 0.08975649df |> map_dbl(median)## a b c d
## -0.2855340 -0.5648310 -0.3025734 -0.2135869df |> map_dbl(sd)## a b c d
## 0.6171634 0.7422437 1.0691040 0.9295846- The second argument,
.f, the function to apply, can be a formula, a character vector, or an integer vector. map_*()uses.feach time it’s called
map_dbl(df, mean, trim = 0.5)## a b c d
## -0.2855340 -0.5648310 -0.3025734 -0.2135869- Look, the map functions also preserve names.
Shortcuts
- There are a few shortcuts that you can use with
.fin order to save a little typing. Imagine you want to fit a linear model to each group in a dataset. The following toy example splits up the mtcars dataset into three pieces (one for each value of cylinder) and fits the same linear model to each piece:
models <- mtcars |>
split(mtcars$cyl) |>
map(function(dat) lm(mpg ~ wt, data = dat))
models## $`4`
##
## Call:
## lm(formula = mpg ~ wt, data = dat)
##
## Coefficients:
## (Intercept) wt
## 39.571 -5.647
##
##
## $`6`
##
## Call:
## lm(formula = mpg ~ wt, data = dat)
##
## Coefficients:
## (Intercept) wt
## 28.41 -2.78
##
##
## $`8`
##
## Call:
## lm(formula = mpg ~ wt, data = dat)
##
## Coefficients:
## (Intercept) wt
## 23.868 -2.192- The syntax for creating an anonymous function in R is quite verbose so purrr provides a convenient shortcut: a one-sided formula
models <- mtcars |>
split(mtcars$cyl) |>
map(~lm(mpg ~ wt, data = .))- Here we have used
.as a pronoun: it refers to the current list element (in the same way that [i] referred to the current index in the for loop).
- When you’re looking at many models, you might want to extract a summary statistic like the
summary()and then extract the component calledr.squared. We could do that using the shorthand for anonymous functions:
models |>
map(summary) |>
map_dbl(~.$r.squared)## 4 6 8
## 0.5086326 0.4645102 0.4229655- Or
models |>
map(summary) |>
map_dbl("r.squared")## 4 6 8
## 0.5086326 0.4645102 0.4229655- You can also use an integer to select elements by position:
x <- list(list(1, 2, 3), list(4, 5, 6), list(7, 8, 9))
x |> map_dbl(2)## [1] 2 5 8Mapping over multiple arguments
- So far we’ve mapped along a single input. But often you have multiple related inputs that you need iterate along in parallel. That’s the job of the
map2()andpmap()functions. For example, imagine you want to simulate some random normals with different means and different SDs.
mu <- list(5, 10, -3)
sigma <- list(1, 5, 10)
map2(mu, sigma, rnorm, n = 5) |> str()## List of 3
## $ : num [1:5] 5.87 5.19 4.66 6.64 3.68
## $ : num [1:5] 12.72 6.33 5.68 4.46 17.07
## $ : num [1:5] 5.29 -11.73 -3.15 1.01 13.79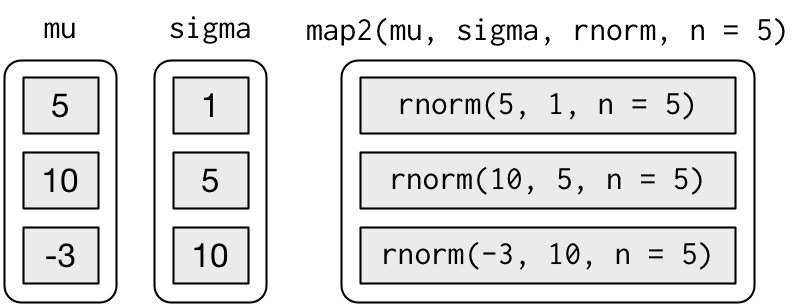
- You could also imagine
map3(),map4(),map5(),map6()etc, but that would get tedious quickly. Instead, {purrr} providespmap()which takes a list of arguments
mu <- list(5, 10, -3)
sigma <- list(1, 5, 10)
n <- list(1, 3, 5)
list(n, mu, sigma) |>
pmap(rnorm) |>
str()## List of 3
## $ : num 5.34
## $ : num [1:3] 4.72 -2.94 17.51
## $ : num [1:5] 0.468 0.181 -11.371 2.34 2.545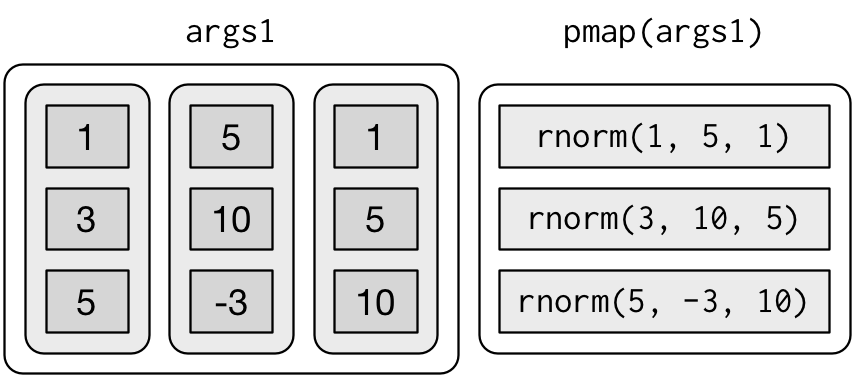
- Since the arguments are all the same length, it makes sense to store them in a data frame:
params <- tribble(
~mean, ~sd, ~n,
5, 1, 1,
10, 5, 3,
-3, 10, 5
)
params %>%
pmap(rnorm)## [[1]]
## [1] 5.182253
##
## [[2]]
## [1] 19.639768 1.119090 8.286335
##
## [[3]]
## [1] -3.222255 10.904046 16.163897 -12.773508 -6.674612